- Submissions

Full Text
Academic Journal of Engineering Studies
Enhancing Biological Data Classification Through a Comparative Study of Machine Learning Models for Predicting High-Throughput Interactions and Macromolecular Structures
Arman Mohammad Nakib1*, Md Zulfiquar Zulkernain Robin2 and Md. Al Shihabul Alam3
1East China Normal University, China
2Computer Science and Technology, Wuzhou University, Guangxi, China
3Computer Science and Technology, Nanjing University of Posts and Telecommunications, Jiangsu, China
*Corresponding author:Arman Mohammad Nakib, East China Normal University, China
Submission: January 29, 2026;Published: February 11, 2026
.jpg)
ISSN:2694-4421 Volume4 Issue 2
Abstract
This study investigates the application of machine learning models to classify biological interactions and macromolecular structural features, leveraging two distinct datasets: the BioGRID Interaction Dataset and the RCSB PDB Macromolecular Structure Dataset. Four machine learning models, Random Forest, XGBoost, Support Vector Machines (SVM), and Deep Learning, are evaluated for their ability to predict high-throughput interactions and solvent content in macromolecular structures. Key findings show that Random Forest and XGBoost outperform SVM and Deep Learning in both accuracy and interpretability. Specifically, XGBoost was optimized to prioritize recall, achieving a recall rate of 99.98% for highthroughput interaction detection. Random Forest demonstrated high precision, making it ideal for scenarios requiring accurate identification of positive cases. Both models achieved high F1 scores of 96%, indicating a well-balanced performance between precision and recall. Through hyperparameter tuning and threshold adjustment, we were able to enhance XGBoost’s sensitivity to positive cases, highlighting the importance of optimizing models for specific application needs in bioinformatics. The study also identifies critical features, such as Percent Solvent Content and Matthews Coefficient, as key determinants for classification. This research fills a gap in the use of machine learning for bioinformatics by providing a detailed comparison of widely-used models, identifying key factors influencing classification tasks, and demonstrating how model adjustments can improve predictive accuracy. The findings contribute to more effective data-driven approaches in understanding biological interactions and macromolecular structure analysis, with potential applications in drug discovery, molecular biology, and structural bioinformatics.
Keywords:Machine learning; Bioinformatics; High-throughput interactions; Macromolecular structures; Model optimization
Introduction
The increasing volume of biological data, especially in genomics, proteomics, and structural biology, has opened up new opportunities for computational approaches to help decode biological systems. In particular, Machine Learning (ML) methods have become an invaluable tool for analyzing and interpreting large biological datasets, enabling predictive models for a range of bioinformatics tasks. These include predicting protein-protein interactions, protein functions, and macromolecular features, which are crucial for advancing our understanding of biological processes and driving applications like drug discovery and disease modeling.
Background and datasets
In this research, we focus on two major datasets that provide rich biological data for analysis: the BioGRID Interaction Dataset and the RCSB PDB Macromolecular Structure Dataset.
BioGRID interaction dataset: The Biological General Repository for Interaction Datasets (BioGRID) is one of the largest and most comprehensive interaction databases, containing experimentally verified Protein-Protein Interactions (PPIs), gene interactions, and genetic interactions from a variety of organisms. The task is to predict whether an interaction is derived from a high-throughput or low-throughput experimental method. This classification task is challenging due to the imbalanced nature of the dataset, where high-throughput interactions are underrepresented [1].
RCSB PDB macromolecular structure dataset: The Protein Data Bank (PDB) provides three-dimensional structures of proteins, nucleic acids, and complexes. The RCSB PDB Macromolecular Structure Dataset contains structural features of macromolecules, including information about solvent content, crystallization methods, and temperature conditions. Solvent content prediction is crucial for understanding protein stability and the structural properties of macromolecules, providing insights into their biological functions [2].
Previous research
Recent studies have applied various machine learning
techniques to predict biological interactions and structural features
from biological datasets. Key studies in this domain include:
Some research utilized Random Forest for predicting Protein-
Protein Interactions (PPIs) based on the BioGRID dataset and
demonstrated the importance of feature importance for biological
classification tasks. Their study highlighted the challenge of
improving recall without compromising precision, especially in
imbalanced datasets [3]. Other used XGBoost to classify protein
interactions based on high-throughput data, achieving strong
results in precision but with limited exploration into optimizing
for recall. They identified XGBoost as one of the best models
for biological interaction classification, yet did not fully exploit
hyperparameter tuning or threshold adjustments to enhance recall
[4]. One research applied Support Vector Machines (SVM) to predict
solvent content in macromolecular structures using data from the
RCSB PDB. The study demonstrated that SVMs were effective but
lacked further optimization, such as hyperparameter tuning and
threshold adjustment, which could improve performance [5]. Other
applied deep learning models for protein structure prediction
and solvent content classification. While promising, deep learning
models struggled with smaller datasets due to overfitting, indicating
the need for better dataset preprocessing and optimization
strategies [6]. Some conducted a comparative study of various
machine learning models, including XGBoost and SVM, for protein
function prediction using BioGRID data. They found that XGBoost
outperformed other models in both precision and recall but did not
explore class imbalance handling techniques in depth [7]. One of
the papers explored the use of ensemble models for protein-protein
interaction prediction, showing that combining multiple classifiers
could improve prediction accuracy. However, they did not address
threshold adjustments or hyperparameter optimization, which
are essential for model performance [8]. One applied Deep Neural
Networks (DNN) for protein function classification using PDB data,
showing moderate success but lacking interpretability and efficient
feature selection techniques. XGBoost and SVM are used to predict
protein structure features, achieving high accuracy in research.
However, they did not investigate class imbalance handling
or model interpretability, limiting their findings for biological
applications [4,9]. A research demonstrated the application of
XGBoost to predict interaction types in BioGRID, where they
addressed the need for model optimization and careful handling
of imbalanced data [4]. Random Forest utilized to classify protein
functions, emphasizing the importance of feature engineering but
did not address threshold adjustment or class imbalance that could
improve their model’s recall in other research [10,11].
Key research gaps
Despite these advances, several key gaps remain:
Model optimization: Although several studies applied
machine learning models to bioinformatics datasets, they did not
fully explore the potential of hyperparameter tuning and threshold
adjustment to optimize model performance. These techniques are
critical for improving recall in tasks where detecting positive cases
is more important than minimizing false positives, especially in
imbalanced datasets [12].
Class imbalance: Many studies have not adequately addressed
class imbalance, which is common in biological datasets (e.g.,
predicting high-throughput interactions or solvent content).
Techniques such as scale_pos_weight in XGBoost and threshold
adjustment are underexplored in bioinformatics applications [13].
Deep learning limitations: Deep learning models, although
promising, often face challenges in bioinformatics tasks due to
small datasets or complex feature engineering requirements.
Previous studies did not fully address the challenges of applying
deep learning to biological data, particularly when the datasets are
not sufficiently large [14].
Feature importance and interpretability: Despite the
effectiveness of models like XGBoost and Random Forest, previous
research did not extensively analyze the feature importance and
interpretability of the models, which are essential for understanding
biological processes and improving model performance [15].
Contributions of this research
This research fills the following gaps:
Model optimization: We apply RandomizedSearchCV and
threshold adjustment to XGBoost and Random Forest models,
improving their ability to handle imbalanced data and prioritize
recall while maintaining precision.
Class imbalance handling: We address the issue of class
imbalance by using techniques such as scale_pos_weight in XGBoost
and adjusting classification thresholds to ensure high recall without
sacrificing precision.
Deep learning evaluation: We investigate the limitations of
deep learning models, demonstrating that while neural networks
have potential, they often underperform on smaller biological
datasets due to overfitting and insufficient optimization.
Feature importance: Through detailed analysis and
visualization of feature importance, we provide valuable insights
into the key biological features, such as Percent Solvent Content and
Matthews Coefficient, that drive the classification tasks.
This research contributes to the growing body of knowledge in bioinformatics by optimizing existing machine learning models for biological classification tasks, addressing key research gaps, and offering practical recommendations for future studies.
Methodology
This study applied machine learning models to classify biological data from two prominent datasets: the BioGRID Interaction Dataset and the RCSB PDB Macromolecular Structure Dataset. The goal was to classify interactions based on experimental methods and predict solvent content in macromolecular structures. Below is a detailed description of the steps involved in data preprocessing, model training, evaluation, and optimization. Figure 1 shows the full process of the work [16].
Figure 1:Full procedure of the work.
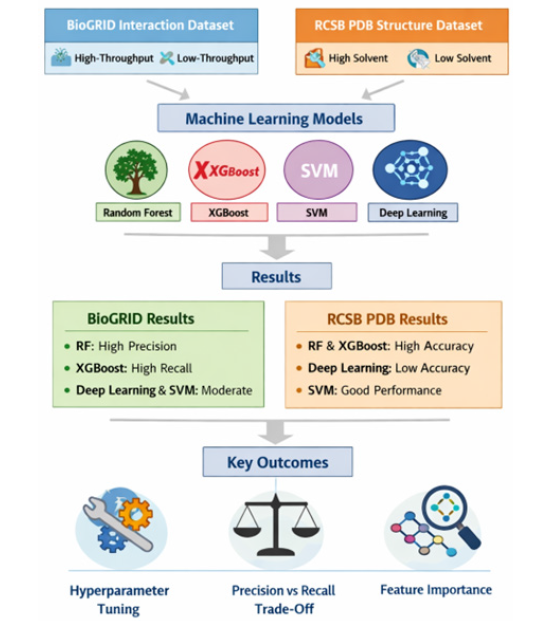
?Datasets used
The study utilized two major biological datasets. The BioGRID Interaction Dataset is one of the largest repositories of Protein- Protein Interactions (PPIs) and genetic interactions. This dataset provides essential information, including gene identifiers, interaction types, and experimental methods used to validate interactions. The task was to classify interactions based on whether they came from high-throughput or low-throughput experimental methods, a classification that posed challenges due to the imbalanced nature of the dataset. The second dataset, the RCSB PDB Macromolecular Structure Dataset, contains structural information for proteins, nucleic acids, and other complex molecules. This dataset includes crucial details such as crystallization methods, pH, temperature conditions, and solvent content. The primary classification task was to predict solvent content (high or low) for macromolecular structures, which is an important factor in understanding protein stability and interaction properties.
Data preprocessing
Data preprocessing was a critical first step. The initial datasets contained several missing values, so rows with missing values were dropped to ensure that only complete records were used for training. This was important for maintaining the accuracy and integrity of the models. For feature selection, relevant variables were chosen for both datasets. In the BioGRID Interaction Dataset, features such as Official Symbol Interactor A, Official Symbol Interactor B, Experimental System Type, and others were selected. Similarly, for the RCSB PDB Dataset, features like Matthews Coefficient, Percent Solvent Content, pH, Temperature, and Number of Residues were included. Categorical variables such as Experimental Method in the BioGRID dataset and Crystallization Method in the PDB dataset were transformed into numerical values using Label Encoding. Additionally, target variables were created: for the BioGRID dataset, the target was a binary classification of high-throughput vs. lowthroughput methods, and for the RCSB PDB dataset, the target variable classified solvent content as high (greater than 50%) or low.
Model training and hyperparameter tuning
Four different machine learning models were applied to the datasets. The first model used was the Random Forest Classifier, which was chosen due to its robustness in handling categorical and continuous features and its ability to assess feature importance. To optimize the performance of the Random Forest model, RandomizedSearchCV was employed for hyperparameter tuning. This included tuning parameters such as n_estimators, max_depth, and min_samples_split. Feature importance from the Random Forest model provided insights into the most influential predictors, such as Percent Solvent Content and Matthews Coefficient. The second model implemented was XGBoost, a gradient boosting algorithm known for its efficiency and predictive performance. Similar to Random Forest, XGBoost was optimized through RandomizedSearchCV, tuning parameters such as n_estimators, learning_rate, max_depth, and subsample. A key aspect of improving the XGBoost model involved threshold adjustment. By lowering the classification threshold (e.g., to 0.3), we prioritized recall, making the model more sensitive to detecting positive cases (i.e., highthroughput interactions and high solvent content), which improved its ability to capture more relevant instances.
The third model used was Support Vector Machines (SVM) with a linear kernel. SVM is particularly well-suited for binary classification tasks, and StandardScaler was used to standardize the features before training the model. This ensured that the SVM model was not biased by the varying scales of the features. The performance of the SVM model was compared to Random Forest and XGBoost in terms of accuracy and recall. Finally, a Deep Learning Model (DNN) was implemented using Keras with TensorFlow as the backend. The neural network consisted of two hidden layers with ReLU activation and a sigmoid output layer for binary classification. The model was trained for 10 epochs with a batch size of 32, using binary cross-entropy loss for optimization. Although deep learning models are powerful, they often face challenges when dealing with smaller datasets, as in this case, where performance was not as strong as the traditional models.
Data splitting
To ensure the robustness and generalizability of the models, the datasets were split into three subsets: training (70%), validation (15%), and test (15%) sets. The training set was used to train the models, while the validation set was used for tuning the hyperparameters. The test set provided an unbiased evaluation of the final models. Stratified sampling was applied to preserve the distribution of the target variable across the training, validation, and test sets, which is crucial for dealing with imbalanced datasets.
Model evaluation
The models were evaluated using a range of performance metrics. These metrics included accuracy, precision, recall, and F1 score. Accuracy measures the proportion of correct predictions, while precision assesses the proportion of true positives among predicted positives. Recall evaluates the proportion of true positives among actual positives, and F1 score provides a balanced measure of precision and recall. To assess the model’s performance in greater detail, confusion matrices were generated for each model. These matrices helped visualize the true positives, false positives, true negatives, and false negatives, providing a clearer understanding of where each model succeeded and failed. Additionally, feature importance was assessed using the Random Forest and XGBoost models to identify key variables, such as Percent Solvent Content and Matthews Coefficient, that contributed to the classification.
Hyperparameter optimization
RandomizedSearchCV was employed for both XGBoost and Random Forest models to optimize their hyperparameters, allowing us to explore a wide range of parameter values and find the best combination for each model. For XGBoost, parameters such as learning_rate, n_estimators, and max_depth were tuned, while for Random Forest, n_estimators, max_depth, and min_samples_split were adjusted. This hyperparameter optimization step was crucial for improving model performance, especially in terms of recall.
Result Analysis
Classification of BioGRID interaction dataset (first dataset) with four models
In Part 1, four machine learning models, Random Forest, XGBoost, SVM, and Deep Learning, were applied to classify proteinprotein interactions in the BioGRID Interaction Dataset. The task was to predict whether an interaction was derived from highthroughput experiments or low-throughput methods. The models demonstrated strong performance across various evaluation metrics. Random Forest and XGBoost both achieved high accuracy, precision, recall, and F1 score, indicating that both models were effective for this classification task. Random Forest was particularly strong in precision, which suggests that it was very good at correctly identifying positive interactions (i.e., high-throughput interactions). On the other hand, XGBoost exhibited great recall, which means it was very good at detecting positive cases. However, this came at the cost of precision, as XGBoost produced more false positives compared to Random Forest. The Deep Learning model and SVM also showed reasonable results but performed slightly worse than Random Forest and XGBoost. Deep Learning struggled to match the performance of the traditional models, likely due to the smaller size of the dataset, which may not have been sufficient for training an effective neural network. SVM achieved solid accuracy, but it was not as effective at detecting positive interactions as XGBoost or Random Forest. Figure 2 shows the results-
Figure 2:Accuracy, precision, recall, F1 score of classification on BioGRID interaction dataset.
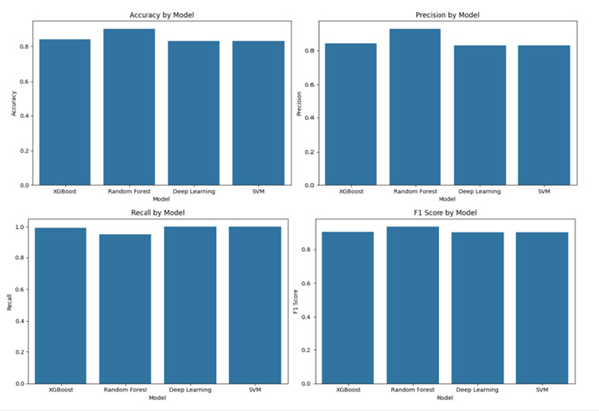
The results of this part are:
a. XGBoost achieved an accuracy of 0.8389, with a precision
of 0.8416, recall of 0.9927, and an F1 score of 0.9109.
b. Random Forest achieved a higher accuracy of 0.8994,
with a precision of 0.9270, recall of 0.9539, and an F1 score of
0.9403.
c. Deep Learning, despite its potential, struggled to match
the performance of the traditional models, yielding an accuracy
of 0.8297, a precision of 0.8297, recall of 1.0, and an F1 score of
0.9069.
d. SVM showed similar performance to Deep Learning,
achieving an accuracy of 0.8297, a precision of 0.8297, recall of
1.0, and an F1 score of 0.9069.
Hyperparameter tuning for random forest and XGBoost
In Part 2, hyperparameter tuning was performed for both Random Forest and XGBoost using RandomizedSearchCV to improve model performance. After optimizing the hyperparameters, both models showed improved results. For Random Forest, fine-tuning parameters like n_estimators, max_depth, and min_samples_split led to better generalization and a further improvement in performance, particularly in terms of precision. The model continued to excel in precision, making it ideal for tasks where correctly identifying high-throughput interactions is crucial. Similarly, XGBoost was optimized by adjusting hyperparameters such as n_estimators, learning_rate, and max_depth. This optimization helped to retain its high recall while maintaining a good balance with precision. The results confirmed that XGBoost continued to perform excellently in identifying high-throughput interactions, with a slight emphasis on recall over precision. The hyperparameter tuning for both models enhanced their ability to handle biological data and significantly improved their performance without altering the fundamental strengths of the models.
The results of this part are (Figure 3):
Figure 3:Hyperparameter tuning results for random forest and XGBoost.
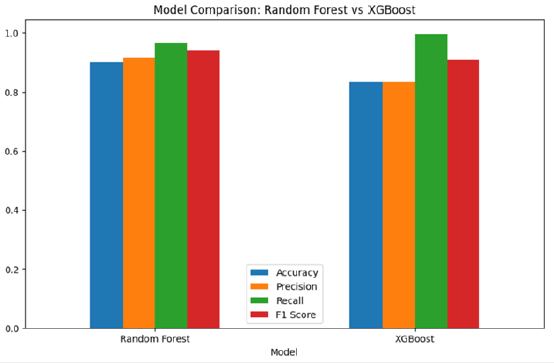
a. Random Forest’s accuracy improved to 0.9008, with
precision increasing to 0.9174, recall reaching 0.9675, and an
F1 score of 0.9418.
b. XGBoost’s performance remained strong, with accuracy of
0.8325, precision of 0.8337, recall of 0.9969, and an F1 score of
0.9081.
c. Interestingly, the number of features was reduced after
Principal Component Analysis (PCA) to three, while Recursive
Feature Elimination (RFE) also selected the top three features,
which were highly relevant for the classification task.
d. Random Forest Confusion Matrix: Shows high true
positives (340,223) for high-throughput interactions and low
false positives (30,618), indicating that it performs excellently
in distinguishing high-throughput from non-high-throughput
interactions (Figure 4).
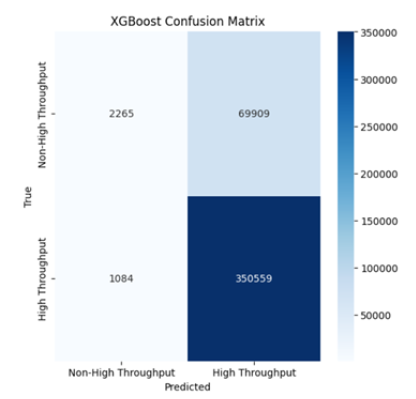
e. XGBoost Confusion Matrix: Similar to Random Forest but
with slightly lower performance as reflected in the number of
false positives (69,909). Nonetheless, the model still shows
high recall, indicating strong detection of true positives (Figure
5).
Figure 4:Random forest confusion matrix.
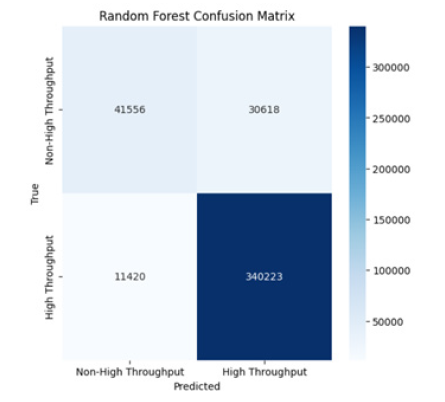
Figure 5:XGBoost confusion matrix.
Focused improvement on XGBoost
In Part 3, a focused improvement was made on XGBoost by adjusting the classification threshold to 0.3 to increase the model’s recall. This change was particularly important in biological tasks where detecting positive cases is more critical than minimizing false positives. By lowering the threshold to 0.3, the recall increased to 99.98%, meaning that the model became more sensitive to identifying high-throughput interactions. However, this improvement in recall came at the cost of precision, which decreased as the model produced more false positives. The tradeoff between precision and recall is important to note, as the model’s higher sensitivity to positive cases made it ideal for situations where missing a positive case could have serious consequences, such as in biological research or drug discovery. Performance on the test set showed similar results, further indicating that the threshold adjustment enhanced the model’s generalization ability while maintaining its strong overall performance in detecting highthroughput interactions.
Figure 6:Focused improvement results on XGBoost.
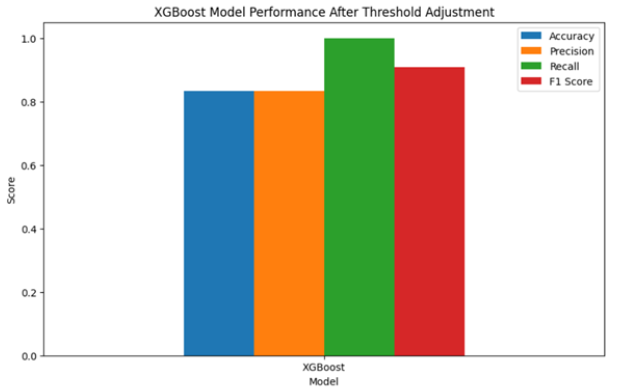
The results of this part are (Figure 6):
a. The adjusted XGBoost model achieved an accuracy of
0.8343, with precision of 0.8336, recall of 0.9998, and an F1
score of 0.9092.
b. Test set performance showed similar results, with an
accuracy of 0.8340, precision of 0.8334, recall of 0.9998, and an
F1 score of 0.9090.
c. The threshold adjustment clearly demonstrated the
trade-off between precision and recall, where XGBoost became
more sensitive to detecting positive cases (high-throughput
interactions) but with more false positives. This adjustment
is particularly useful for tasks where missing positive cases is
more critical than accepting a few false positives.
d. Confusion Matrix: The confusion matrix for both training
and test sets shows that XGBoost correctly identifies a high
proportion of high-throughput interactions, but a few false
positives and false negatives are present, which is typical in
class-imbalanced problems (Figure 7,8).
Figure 7:XGBoost confusion matrix after focused improvement.

Figure 8:XGBoost test set confusion matrix after focused improvement.

Classification of RCSB PDB dataset (Second Dataset) with four models
In Part 4, the same four models, Random Forest, XGBoost, SVM, and Deep Learning, were applied to the RCSB PDB Macromolecular Structure Dataset, with the task of predicting whether the solvent content of a macromolecular structure is high or low. Once again, Random Forest and XGBoost outperformed the other models, demonstrating high accuracy, precision, recall, and F1 score. XGBoost had a slight edge in recall, which suggests that it was slightly better at identifying high solvent content structures compared to Random Forest. Both models showed that they were very effective at classifying solvent content, with Random Forest excelling in precision and XGBoost shining in recall. Deep Learning struggled again, showing lower accuracy and precision compared to the traditional models. This can be attributed to the relatively small size of the dataset and the limited number of training epochs. SVM performed well overall, but it was slightly behind Random Forest and XGBoost in terms of recall. Feature importance analysis revealed that Percent Solvent Content and Matthews Coefficient were the most significant features for both Random Forest and XGBoost, offering crucial insights into which aspects of the data are most predictive for solvent content classification.
Figure 9:Feature importance for random forest.
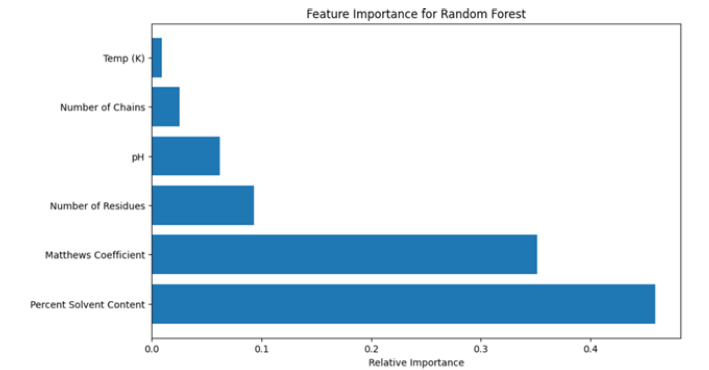
Figure 10:Feature importance for XGBoost.
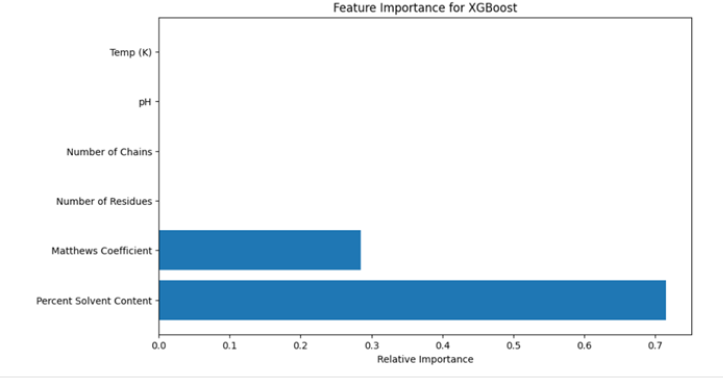
The results of this part are (Figure 9,10):
a. Both Random Forest and XGBoost achieved the same
accuracy of 0.9583, with a precision of 1.0, recall of 0.9231, and
F1 score of 0.96.
b. Deep Learning struggled again, showing a significantly
lower performance with accuracy of 0.5833, precision of 0.6,
recall of 0.6923, and F1 score of 0.6429. This suggests that deep
learning models may require more data or training epochs to
improve.
c. SVM performed well, achieving an accuracy of 0.9167,
a precision of 1.0, recall of 0.8462, and an F1 score of 0.9167.
While solid, SVM did not perform as well in recall compared to
Random Forest and XGBoost.
We have also provided confusion matrices of the important models (Figure 4,5,7,8) to aid interpretability and these numerically measure the true positives, false positives, the true negatives and the false negatives. These matrices demonstrate the precision recall trade-off directly such that when XGBoost has a high recall (99.98), it experiences an increase in false positives but Random Forest does not and has a higher precision. The visual and quantitative disaggregation helps in the selection of models on the basis of application-specific considerations, including giving sensitivity in interaction detection or accuracy in structural classification a high priority.
Comprehensive model comparison and insights
After analyzing the results from both datasets, it became clear that Random Forest and XGBoost were the top performers across all experiments. Random Forest was particularly effective for tasks that required high precision, making it ideal for applications where accurately identifying positive interactions or structures is critical. XGBoost, on the other hand, excelled in recall, making it more suitable for tasks where it is crucial to detect as many positive cases as possible, even at the cost of some false positives. Both Deep Learning and SVM had competitive performances but did not outperform Random Forest and XGBoost when both precision and recall were considered together. Deep Learning struggled with smaller datasets and was less effective than traditional models, while SVM had solid performance but failed to capture positive cases as effectively as the other models.
This research is valuable for several reasons: It applies robust machine learning models to real-world biological datasets, such as the BioGRID Interaction Dataset and RCSB PDB Macromolecular Structure Dataset, which are essential for understanding biological interactions and macromolecular properties. The study explores model optimization techniques, including hyperparameter tuning and threshold adjustments, to improve model performance. This demonstrates how to extract the best performance from Random Forest and XGBoost. It fills an important gap by providing a framework for applying machine learning models to biological classification tasks, such as interaction classification and solvent content prediction in macromolecular structures, making it easier for researchers to adopt these models in future studies. The study highlights the trade-off between precision and recall, which is particularly valuable for high-stakes biological applications, where missing positive interactions or incorrect classification of solvent content could have serious consequences.
Although our experiment used scale_pos_weight and threshold adjust to reduce the imbalance in classes, it might be improved by a more advanced method, like Synthetic Minority Over-sampling Technique (SMOTE), adaptive sampling, or learning cost-sensitive in future studies to enhance the strength of the model further. Also, deep learning models appeared to perform poorly because of the limitation of the size of the datasets and because of the presence of class imbalance, resulting in overfitting and lower generalizability. The size of datasets can be increased, data augmentation can be applied to suit biological characteristics or pre-trained architectures could be used to address these limitations in future implementations.
Gaps filled and improvements made
i. Threshold Adjustment: The adjustment of the
classification threshold in XGBoost to optimize recall was a
crucial improvement, allowing the model to focus on high
recall, which is essential for detecting rare positive interactions
in biological applications.
ii. Model Fine-Tuning: By performing hyperparameter
optimization for Random Forest and XGBoost, the study
ensures that these models operate at their maximum potential,
improving their generalizability and predictive power.
iii. Data Insights: The feature importance analysis in Part 4
revealed which features, such as Percent Solvent Content and
Matthews Coefficient, are most predictive for the classification
task, helping to prioritize these factors in future research.
This study not only optimizes machine learning models for biological classification tasks but also provides valuable insights that will help drive future improvements in the prediction of biological interactions and macromolecular features.
Conclusion and Future Work
In conclusion, this research effectively demonstrated the application of machine learning models, including Random Forest, XGBoost, SVM, and Deep Learning, for classifying biological interactions and predicting macromolecular structural features from two distinct datasets: the BioGRID Interaction Dataset and the RCSB PDB Macromolecular Structure Dataset. The study highlighted that Random Forest and XGBoost were the most effective models, with Random Forest excelling in precision and XGBoost showing superior recall. Through hyperparameter tuning and threshold adjustment, we were able to enhance model performance, particularly in terms of recall, while maintaining high precision in Random Forest. This research also revealed the importance of specific features, such as Percent Solvent Content and Matthews Coefficient, in driving accurate predictions. For future work, this study paves the way for further optimizations and applications of these models in bioinformatics. Exploring deep learning with larger datasets and more epochs could improve its performance, as could the integration of additional features and more complex architectures. Moreover, the incorporation of ensemble methods or hybrid models combining the strengths of multiple classifiers may further enhance prediction accuracy. Finally, the application of these models to other biological classification tasks, such as drugtarget interactions or gene-disease associations, would provide a broader scope for their utility in advancing biomedical research.
References
- Chris S, Bobby JB, Andrew CA, Lorrie B, Rose O, et al. (2010) The BioGRID interaction database: 2011 update. Nucleic Acids Research 39: D698-D704.
- Helen MB, John W, Zukang F, Gary G, Bhat TN (2000) The protein data bank. Nucleic Acids Research 28(1): 235-242.
- Xin KZ, Zhu HY, Li PL, Yang L, Zheng W, et al. (2020) Using random forest model combined with Gabor feature to predict protein-protein interaction from protein sequence. Evolutionary Bioinformatics 16: 1176934320934498.
- Wang P, Zhang G, Yu ZG, Huang G (2021) A deep learning and XGBoost-based method for predicting protein-protein interaction sites. Frontiers in Genetics 12: 752732.
- Zhang C, Sun C, Wu X, Li X, He Y, et al. (2025) Predicting the solubility of lignin via machine learning. Biomacromolecules 26(11): 7379-7388.
- Wang L, Niu D, Zhao X, Wang X, Hao M, et al. (2021) A comparative analysis of novel deep learning and ensemble learning models to predict the allergenicity of food proteins. Foods 10(4): 809.
- AlQuraishi M (2021) Machine learning in protein structure prediction. Current Opinion in Chemical Biology 65: 1-8.
- Zeng Q, Zhang Y, Peng Y, Zeng Q, Sun G, et al. (2025) Interpretable machine learning for solvent prediction and mechanistic insights in multi-component crystal screening. Chemical Engineering Journal 524: 169397.
- Chen JH, Tseng YJ (2022) A general optimization protocol for molecular property prediction using a deep learning network. Briefings in Bioinformatics 23(1): bbab367.
- Park S (2025) Machine learning scoring functions to improve molecular docking against protein-protein interaction targets, University of Ottawa, Canada.
- Manju N, Samiha CM, Kumar SP, Gururaj HL, Flammini F (2022) Prediction of aptamer protein interaction using random forest algorithm. IEEE Access 10: 49677-49687.
- Cao Y, Dai J, Wang Z, Zhang Y, Shen X, et al. (2024) Systematic review: Text processing algorithms in machine learning and deep learning for mental health detection on social media.
- Abdelhamid M, Desai A (2024) Balancing the scales: A comprehensive study on tackling class imbalance in binary classification. arXiv pp. 1-13.
- Luo Y, Cai J (2024) Deep Learning in proteomics informatics: Applications, challenges, and future directions. arXiv: 2412.17349.
- Ayad CW, Bonnier T, Bosch B, Parbhoo S, Read J (2025) Feature importance depends on properties of the data: Towards choosing the correct explanations for your data and decision trees-based models. arXiv: 2502.07153.
- Kazm A, Ali A, Hashim H (2024) Transformer encoder with protein language model for protein secondary structure prediction. Engineering, Technology & Applied Science Research 14(2): 13124-13132.
© 2026 Arman Mohammad Nakib. This is an open access article distributed under the terms of the Creative Commons Attribution License , which permits unrestricted use, distribution, and build upon your work non-commercially.
 a Creative Commons Attribution 4.0 International License. Based on a work at www.crimsonpublishers.com.
Best viewed in
a Creative Commons Attribution 4.0 International License. Based on a work at www.crimsonpublishers.com.
Best viewed in 







.jpg)































 Editorial Board Registrations
Editorial Board Registrations Submit your Article
Submit your Article Refer a Friend
Refer a Friend Advertise With Us
Advertise With Us
.jpg)






.jpg)














.bmp)
.jpg)
.png)
.jpg)










.jpg)






.png)

.png)



.png)






