- Submissions

Full Text
Significances of Bioengineering & Biosciences
The Emerging Use of SPH In Biomedical Applications
Milan Toma*
Department of Mechanical Engineering, School of Engineering & Computing Sciences, New York Institute of Technology, USA
*Corresponding author: Milan Toma, Department of Mechanical Engineering, School of Engineering & Computing Sciences, New York Institute of Technology, Old Westbury Campus - HSH116A, Northern Boulevard, Old Westbury, New York 11568-8000, USA
Submission: August 22, 201;Published: October 04, 2017

ISSN 2637-8078Volume1 Issue1
Abstract
Smoothed-particle hydrodynamics is a fast, parallel, modular and low-memory method originally developed for astrophysical applications in three dimensions. In recent years, it has been used for 3D biomedical applications as well. It has proved to be useful in the field of biomedical engineering mostly because the geometries used need to be often patient-specific and therefore complex. Complicated geometries are formed with high number of finite elements which subsequently requires the use of specialized numerical methods. Moreover, computational fluid dynamics in biomedical applications also deals with moving and deforming structures and therefore the use of fluid-structure interaction analyses is important. The utilization of smoothed-particle hydrodynamics in the fluid-structure interaction analysis is ideal to satisfy the need for fast computational method with the use of complicated geometries.
Keywords: Fluid-structure interaction; Smooth particle hydrodynamics; Computer Simulations; Finite Element Method
Introduction
The smoothed-particle hydrodynamics (SPH) method was originally developed to solve astrophysics problems [1] and later used to avoid extreme mesh deformations problems in finite element methods. The lack of a computational grid does affect the accuracy of SPH simulations compared to finite element methods. However, the SPH used in the biomedical applications usually represents the fluid domain, i.e. biofluids in human or animal body interacting with both the biological and mechanical structures. Biological structures are those that are naturally present in the body and interact with biofluids, e.g. cerebrospinal fluid interacting with brain or blood flow interacting with heart valves and blood vessels.
Mechanical structures in human body are sometimes present too, especially when a medical device is implanted to treat a patient with diseased/malfunctioning biological structure, e.g. mechanical heart valves. That kind of numerical simulations requires the use of complicated (patient specific) geometries which then often makes the use of classical finite element methods unsuitable. The number of degrees of freedom is too large and the parallelization of fully coupled fluid-structure interaction (FSI) problems is extremely challenging [2]. However, the SPH interacting with the fully integrated solid finite elements, thus removing the possibility of hourglass modes and element inversion that plagues the classic under-integrated finite element models, yields the ability to reach large (physiologically equivalent) deformations in a timely manner using a standard workstation with graphics processing unit (GPU) technology. And, needless to say, in surgery planning, time can be of the essence; while the cost of the GPU technology is well below the cost of computer clusters.
The need for computational simulations with the capability to provide fast and reliable results to guide the decisions made by the clinicians is axiomatic. For example, single ventricle congenital heart defects afflict 2 per every 1000 births and the success of the surgical procedures designed to correct them can often be questionable [3].
Figure 1: An extra-cardiac total cavopulmonary connection connecting the superior (SVC) and inferior (IVC) venae cavae directly to the pulmonary arteries, bypassing the single ventricle.

Figure 1 shows results of the Fontan procedure which involves redirecting oxygen-poor blood from the top of the body to the lungs by redirecting the blood from the inferior venacava (IVC) to the lungs. At this point, the oxygen-poor blood from upper and lower body flows through the lungs without being pumped. The location of the conduit in relation to the superior vena cava (SVC) and its success at properly redirecting blood flow is largely a hydrodynamics problem. When the conduit is not positioned correctly, insufficient blood flow is redirected through lungs and this extremely invasive surgical procedure can be ineffective [3]. Thus, computer simulations have been used to guide the clinicians’ decisions where to place the conduit [4]. Obviously, when a medical decision is to be made, how much expertise and time it requires to create the 3D models and to run the simulations determine the suitability of the numerical method used and, consequently, how much potential it has to be accepted by the medical community when used in practice.
The SPH has recently been used in modeling the interaction of the blood flow with the arterial walls, heart ventricles and native and artificial heart valves [5-9]. Simulating the interaction of the cerebrospinal fluid with the brain and skull structures using the SPH has also begun [10].
Smoothed Particle Hydrodynamics Formulation
For completeness, the basic governing equations of the SPH method are summarized in this section. The SPH is governed by a set of ordinary differential equations as follows.
The continuity equation, i.e. conservation of mass, is

Where W is the Kernel function and the 4th order diffusive term improves the pressure evolution.
The momentum equation, i.e. Euler equation, is
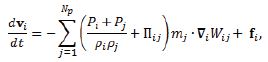
Where IIij is the artificial viscosity.
The energy equation, i.e. first law of thermodynamics, is
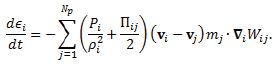
More detail on the numerical methods used in the cardiovascular FSI simulations, e.g. the integration method, artificial viscous term, laminar viscous term, speed of sound used, can be found published elsewhere [5,11].
The SPH interacts with the structural finite elements using a penalty-based contact algorithm. In penalty-based contact, when a penetration is found, a force proportional to the penetration depth is applied to resist and eliminate penetration. Linear contact spring stiffness is based on the nodal masses that come into contact and the time step size as follows
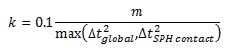
The resulting contact stiffness is independent of the material constants so it is well suited for treating contact between fluid and structure.
Results and Validation
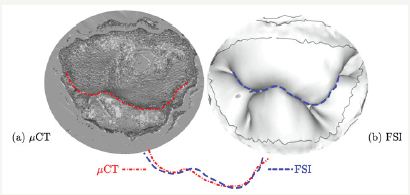
In [7], the SPH was used to simulate and validate the resulting closure of a patient-specific model of mitral valve with detailed 3D chordal structure. The Figure 2 shows the comparison of the closed leaflets reconstructed from CT images with the results of FSI-SPH simulations. Good agreement can be observed.
Figure 2: Closed leaflets reconstructed from CT images compared to the results of FSI simulations [7]. The curves represent the coaptation line where the posterior and anterior leaflets are in contact.
Similarly, in [12] the use of SPH with a comprehensive model of mitral valve was validated against experimental data of papillary muscle forces in its closed configuration during ventricular systole. Once extensively validated, this model can be used to understand varying pathologies, such as the rupture of mitral chordae tendineae [13].
Figure 3: The areas measured across IVC and SVC compared with the MRI measurements
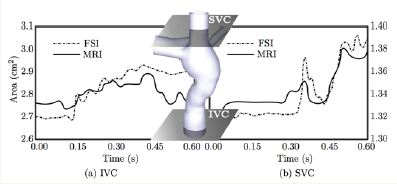
In Figure 3, The IVC and SVC cross-sectional areas are compared with the in vivo MRI measurements [14,15]. Similarities in magnitudes and patterns can be observed.
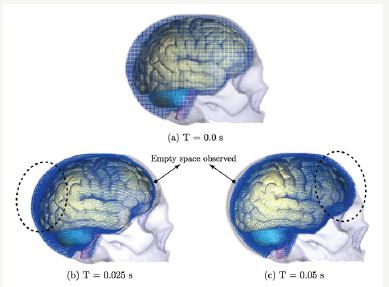
As presented in [10], the use of SPH has been shown the ability to produce results involving all the biofluids surrounding complex structures in the human body, such as cerebrospinal fluid filling the space between the skull and brain. The cushioning effect of cerebrospinal fluid is demonstrated in Figure 4a & 4b shows that during the rapid acceleration phase, fluid particles flow backward, preventing the brain from impacting the occipital/parietal bone and creating an empty space between the anterior brain and frontal bone. During the rapid deceleration phase, fluid particles flow forward, preventing the brain from impacting the frontal bone and creating an empty space between the posterior brain and the occipital/parietal bones, Fig. 4(c). The change from acceleration to deceleration causes the fluid particles to reverse direction. Because the fluid particles move faster than the brain, during the deceleration phase the fluid particles fill in the empty space created in the acceleration phase.
Figure 4: The fluid particles are visibly concentrated in the back of the head at the end of the rapid acceleration phase (a) and at the front of the head at the end of the rapid deceleration phase. (b) The dashed ellipsoids denote the areas where the particle concentration can be observed. The arrows point to the spaces where an empty space is observed after the deformation of the brain and fluid particles migrating with the acceleration/ deceleration.
Discussion
Despite the fact that SPH is not commonly used for biomedical applications, it has been shown that, in combination with FSI, it can provide results that are not only in agreement with the in vivo and in vitro measurements, but can also be obtained in a timely manner, i.e. up to 24 hours on newest GPU technology. The runtime of 5 to 24 hours, depending on the complexity of the model, is significant improvement compared to days and weeks necessary for standard finite element methods to complete even on multiple processors while dealing with ill-conditioned systems and large deformations. And, needless to say, in surgery planning, time can be of the essence. Also, the cost of the GPU technology is well below the cost of computer clusters. Computer simulations have already been used to help the clinicians make their medical decisions. And, when a medical decision is to be made, how much expertise and time it requires to create the 3D models and to run the simulations determine the suitability of the numerical method used. Consequently, it also determines how much potential the numerical method has to be accepted by the medical community when used in practice. As of now, more commonly, it is the academic community that contributes to the decision making process by providing computational results using the patient specific geometries. However, ultimately, if this is to be adapted as routinely used procedure, the academia will have to be omitted from the process. And that can only happen if the computational results are attainable without too much expertise and specialized computer equipment required. The use FSI with SPH is a step forward to achieve this goal. Furthermore, the demand for computational simulations in the medical community has to rise to motivate faster development of such methods. Meantime, while 45 percent of graduating medical students in the USA aspire to work in the academia, only about 16 percent will actually do so and of those who do work in academic settings up to 38 percent will leave academia within 10 years [16].
Though SPH simulations show tremendous promise to be the numerical method of choice when there is the need for timely results on relatively cheap GPU technology, SPH-FSI is not yet fully adapted for the use in biomedical applications. Further development is necessary to readily obtain parameters otherwise commonly used in cardiovascular fluid mechanics, such as powerloss, wall shear stress, and energy dissipation rates. If these features are implemented, SPH-FSI simulations may prove to be a superior tool for biomedical applications.
References
- Rosswog S (2009) Astrophysical smooth particle hydrodynamics. New Astronomy Reviews 53(4-6): 78-104.
- Toma M, Oshima M, Takagi S (2016) Decomposition and parallelization of strongly coupled fluid-structure interaction linear subsystems based on the Q1/P0 discretiza-tion. Computers & Structures 173: 84-94.
- Mondésert B, Marcotte F, Mongeon FP, Dore A, Mercier LA, et al. (2013) Fontan circulation: Success or failure? Can J Cardiol 29(7) 811-820.
- Tree M, Wei ZA, Munz B, Maher K, Deshpande S (2017) A method for in vitro TCPC compliance verification. J Biomech Eng 139(6).
- Jakob Durrwachter (2016) Hemodynamics of the left ventricle: Validation of a smoothed-particle hydrodynamics fluid-structure interaction model. MS thesis, Georgia Insti-tute of Technology, USA.
- Caballero A, Mao W, Liang L, Oshinski J, Primiano C, et al. (2017) Modeling left ventricular blood flow using smoothed particle hydrodynamics. Cardiovascular Engineering and Technology, pp. 1-15.
- Toma M, Bloodworth CH, Einstein DR, Pierce EL, Cochran RP (2016) High resolution subject-specific mitral valve imaging and modeling: Experimental & computational methods. Biomech Model Mechanobiol 15(6): 1619-1630./a>
- Mao W, Li K, Sun W (2016) Fluid-structure interaction study of transcatheter aortic valve dynamics using smoothed particle hydrodynamics. Cardiovasc Eng Technol 7(4): 374-388.
- Mao W, Li K, Caballero A, Sun W (2016) Fluid–structure interaction study of transcatheter aortic valve dynamics using smoothed particle hydrodynamics. Cardiology 134(2).
- Toma M, Nguyen P (2017) Fluid-structure interaction analysis of cerebral spinal fluid with a comprehensive head model subject to a car crash-related whiplash. 5th International Conference on Computational and Mathematical Biomedical Engineering - CMBE2017.
- Toma M, Einstein DR, Bloodworth CH, Cochran RP, Yoganathan AP (2017) Fluid-structure interaction and structural analyses using a comprehensive mitral valve model with 3D chordal structure. Int J Numer Method Biomed Eng 33(4).
- Toma M, Jensen MØ, Einstein DR, Yoganathan AP, Cochran RP, et al. (2016) Fluid-structure interaction analysis of papillary muscle forces using a comprehensive mitral valve model with 3D chordal structure. Ann Biomed Eng 44(4): 942-953.
- Toma M, Bloodworth CH, Pierce EL, Einstein DR, Cochran RP (2017) Fluid-structure interaction analysis of ruptured mitral chordae tendineae. Ann Biomed Eng 45(3): 619-631.
- Frakes D, Smith M, de Zélicourt D, Pekkan K, Yoganathan A (2004) Threedimensional velocity field reconstruction. J Biomech Eng 126(6): 727- 735.
- Sundareswaran KS, Frakes DH, Fogel MA, Soerensen DD, Oshinski JN, et al. (2009) Optimum fuzzy filters for phase-contrast magnetic resonance imaging segmentation. J Magn Reson Imaging 29(1): 155-165.
- AAMC (2015) Medical school graduation questionnaire. Tech Rep Association of American Medical Colleges.
© 2017 Milan Toma. This is an open access article distributed under the terms of the Creative Commons Attribution License , which permits unrestricted use, distribution, and build upon your work non-commercially.
 a Creative Commons Attribution 4.0 International License. Based on a work at www.crimsonpublishers.com.
Best viewed in
a Creative Commons Attribution 4.0 International License. Based on a work at www.crimsonpublishers.com.
Best viewed in 







.jpg)































 Editorial Board Registrations
Editorial Board Registrations Submit your Article
Submit your Article Refer a Friend
Refer a Friend Advertise With Us
Advertise With Us
.jpg)






.jpg)














.bmp)
.jpg)
.png)
.jpg)










.jpg)






.png)

.png)



.png)






