- Submissions

Full Text
Research in Medical & Engineering Sciences
Study of Comparison of Oral Microbial Flora of Smokers vs Non-Smokers in a Metropolitan City
Shyam Sundar Tiwari1, Atul Raj2*, Sayan Bhattacharyya2* and Abhirup Ganguli3
1M.Sc. Biotechnology student, Swami Vivekanand Institute of Modern Science, Sonarpur, India
2Department of Microbiology, All India Institute of Hygiene and Public Health, India
3Department of Microbiology, Swami Vivekananda Institute of Modern Science, Sonarpur, India
*Corresponding author:Atul Raj and Sayan Bhattacharyya, Department of Microbiology, All India Institute of Hygiene and Public Health, Kolkata, India
Submission: September 19, 2023;Published: October 20, 2023

ISSN: 2576-8816Volume10 Issue4
Abstract
There are more than 700 different bacteria found in oral cavity. Epidemiological studies show significantly higher risk for periodontal disease in smokers compared to non-smokers. The increased risk is proportional to the duration and rate of smoking. This is an important area of public health research. In our study we found that MRSA was significantly more in oral cavity of smokers when compared to non-smokers.
Keywords:Oral Microbiome; Pathogenic microorganisms; Smokers; Nonsmokers
Abbreviations:MRSA: Methicillin Resistant Staphylococcus aureus
Introduction
Oral microbiome
The inside of our mouth is the perfect place for bacteria to thrive: it’s dark, it’s warm, it’s wet and the foods and drinks we consume provide nutrients for them to eat. But when the harmful bacteria build up around our teeth and gums, we are at risk of developing periodontal (or gum) disease, experts say, which is an infection and inflammation in the gums and bone that surround your teeth. Such conditions in your mouth may influence the rest of your body. A growing yet limited body of research, for instance, has found that periodontal disease is associated with a range of health conditions including diabetes, heart disease, respiratory infections and dementia. There are more than 700 bacteria found in the oral cavity ,2nd highest after gut.
The community of microbial residents in our body is called the microbiome. The term “microbiome” is coined by Nobel Prize laureate Joshua Lederberg, to describe the ecological community of symbiotic, commensal and pathogenic microorganisms. These microorganisms literally share our body space. Oral microbiome was first identified by the Dutchman Antony van Leeuwenhoek who first identified oral microbiome using a microscope constructed by him. He was called the father of microbiology and a pioneer who discovered both protists and bacteria. In 1674, he observed his own dental plaque and reported “little living animalcules prettily moving.” Collective genome of microorganisms that reside in the oral cavity is called oral microbiome. In periodontal disease, tooth loss, and possibly gastrointestinal cancers oral bacteria playing a critical role and several physiological functions. The bacterial community structure of the oral microbiome is formed principally by Firmicutes, Bacteroidetes, Proteobacteria, Actinobacteria, Spirochaetes, and Fusobacteria, with only 4% of species belonging to other phyla. The number of microbes present in our bodies is almost the same or even more [1].
Oral microbiome and disease
The oral microbiome is known to participate in metabolic functions that include the deglycosylation of complex carbohydrates by Streptococcus oralis? (Byers et al., 1999) and sulfate reduction by Desulfobacter and Desulfovibrio (van der Hoeven et al., 1995). Proteolytic activities of periodontal bacteria also are important for amino acid absorption. Proteolysis is mostly produced by Porphyromonas, Prevotella, and Fusobacterium species (Jie Bao et al., 2008), and an aminopeptidase effect has been observed from Fusobacterium nucleatum? (Rogers et al., 1998) [2].
The oral microbiota contains one of the highest diversities of bacteria in the human body. Many of these bacteria are believed to contribute to the development of cancer. Fusobacterium nucleatum (F. nucleatum) is one such bacteria and is considered an oncobacterium. F. nucleatum is a Gram-negative anaerobic bacterium that is found primarily in the oral mucosa under normal conditions. However, F. nucleatum has been implicated in several disease states including colorectal cancer. F. nucleatum is more abundantly found in diseased lesions. Multiple studies have shown that F. nucleatum is found to be present in potentially high numbers in cancerous tissues [3].
The stability of the oral microbiome may be important for resistance against colonization by pathogens (Vollaard and Clasener, 1994) and is mainly produced by the presence of commensal streptococci, which produce a high amount of bacteriocins and inhibit the growth of Gram negatives. Streptococcus salivarius has been identified as a member of the normal oral microbiome that may maintain the bacterial community structure and has been suggested as a possible probiotic.
Smoking has been recognized as a major risk factor for periodontal disease, affecting the prevalence, severity, progression and treatment response of the disease, second only to the dental plaque. Epidemiological studies have presented a significantly higher risk for periodontal disease in smokers compared to nonsmokers and the increased risk is proportional to the duration and rate of smoking. Cigarette smoking is a public health problem. It decreases the commensal population of normal flora in the oral cavity leading to an increase of pathogenic microbes [4].
Active smokers and those exposed to second hand smoke are at increased risk of bacterial infection. Tobacco smoke exposure increases susceptibility to respiratory tract infections, including tuberculosis, pneumonia and Legionnaires disease; bacterial vaginosis and sexually transmitted diseases, such as chlamydia and gonorrhoea; Helicobacter pylori infection; periodontitis; meningitis; otitis media; and post-surgical and nosocomial infections. Tobacco smoke compromises the anti-bacterial function of leukocytes, including neutrophils, monocytes, T cells and B cells, providing a mechanistic explanation for increased infection risk. Further epidemiological, clinical and mechanistic research into this important area is warranted [5].
Gingivitis
It is not uncommon to have some form of periodontal disease. In its early stages, called gingivitis, the gums may become swollen, red or tender and may bleed easily. If left untreated, gingivitis may escalate to periodontitis, a more serious form of the disease where gums can recede, bone can be lost, and teeth may become loose or even fall out. With periodontitis, bacteria and their toxic byproducts can move from the surface of the gums and teeth and into the bloodstream, where they can spread to different organs, said Ananda P. Dasanayake, a professor of epidemiology at the New York University College of Dentistry, US. This can happen during a dental cleaning or flossing, or if you have a cut or wound inside your mouth, he said.
If you have inflammation in the mouth that is untreated, some of the proteins responsible for that inflammation can spread throughout the body and potentially damage other organs.
Diabetes
Of all the associations between oral health and disease, the one with the most evidence is between periodontal disease and diabetes. And the two conditions seem to have a two-way relationship. periodontal disease seems to in-crease the risk for diabetes, and vice versa. Researchers have yet to understand exactly how this might work, but in one review published in 2017, researchers wrote that the systemic inflammation caused by periodontal disease could worsen the body’s ability to signal for and respond to insulin, leading to diabetes
In another study, published in April 2023, scientists found that diabetics who were treated for periodontal disease saw their overall healthcare costs decrease by 12 to 14 per cent. “You treat periodontal disease, you improve the diabetes,” Dasanayake said.
Pneumonia
If large amounts of bacteria from the mouth are inhaled and settle in the lungs, that can result in bacterial aspirations pneumonia. This phenomenon has been observed mainly in patient who are hospitalized or older adults in nursing homes and is a concern for those who can’t floss or brush their teeth on their own.
Cardiovascular diseases/
In a report published in 2020, an international team of experts concluded that there is a significant link between periodontitis and increased chances of heart attack stroke, plaque buildup in the arteries and other cardiovascular conditions. A 2012 statement from the American heart association noted that inflammation in the gums has been associated with higher levels of inflammatory proteins in blood that have been linked with poor heart disease [6].
Pregnancy complications
A number of studies and reviews have found association between severe periodontal disease and preterm, low birth weight babies. Though more research is needed to confirm this link.
In 2019 reviews, researchers found that treating periodontal disease during pregnancy improve birth weight and reduced the risk of preterm birth and the death of foetus or the newborn. And in 2009 study researchers found that oral bacteria could travel to the placenta-potentially playing a role in chorloamnionitis a serious infection of the placenta and amniotic fluid that can lead to an early delivery or even cause life threatening complications if left untreated [7].
Cystic fibrosis
Cystic fibrosis is a life-shortening autosomal recessive disease affecting the pulmonary, gastrointestinal and genitourinary systems. Lung colonization with a succession of pathogens, such as Staphylococcus aureus and H. influenzae is likely to occur in affected individuals. Ultimately, persistent colonization by the opportunistic pathogen Pseudomonas aeruginosa contributes to significant morbidity and mortality in cystic fibrosis [5-8]. Epidemiological studies have shown that second hand smoke exposure is associated with poor prognosis in cystic fibrosis patients [9] and that a dosedependent relationship exists between number of cigarettes smoked and severity of disease amongst young patients [7,8,10]. In keeping with these data, mice infected with P. aeruginosa and exposed to tobacco smoke exhibit delayed clearance of infection and increased morbidity compared to control mice which were infected but not exposed to smoke [8,9].
Bacterial meningitis
Carriage of H. influenzae, pneumococcus, and meningococcus has been shown to be more common in both active and secondhand smokers than in nonsmokers (34-38). Indeed, the relationship may be dose-dependent, and, in a study of military recruits, smoke exposure gave an attributable risk for meningococcal carriage of 33% (38). In addition to increased carriage, meningitis cases have a two-to fourfold higher risk of exposure to cigarette smoke than controls [10-12]. Furthermore, second-hand smoke exposure predisposes to nasopharyngeal colonization with specific Staphylococcus aureus variants that may have altered capacity to compete with pneumococci subtypes. Passive exposure to tobacco smoke has also been associated with Haemophilus influenzae and pneumococcus meningitis in Australian children [13-15].
Aim of study was
The aim of this study was to find out the difference in oral microbial flora between smokers and Non-Smokers.
Objectives of study were
A. Identification of Microbes.
B. Comparison of microbial flora from the sample of Smokers
and Nonsmokers.
Sample collected from volunteer Before taking swab, written, informed and voluntarily consent was taken from the participants in English, Bengali and Hindi.
Materials and Methods
Time of study
The study was carried out from February 2023 to May 2023.
Place of study
Department of Microbiology, BN campus, All India Institute of Hygiene and Public Health, Kolkata.
Type of study
Laboratory based observational study: 2.4 Fifty-four (54) subjects, representing both genders, ranging in age from 18 to 60. Fifty-four (54) samples were tested. This sample size had been calculated by method of convenience. In case of smokers the swabs were taken from those, who smokes from minimum 5 years. There were 27 swabs taken from smokers and 27 swabs from non-smokers. Two swabs were taken from each volunteer, one for culture and another for Gram stain and Albert’s stain. Gram stain for seeing gram positive and gram-negative bacteria while Albert stain for seeing bacteria containing Metachromatic Granules (MCG) [16,17].
Methodology proper
These volunteers are from
A. Bidhannagar
B. Urban health unit, All India Institute of Hygiene and Public
health, Chetla
C. Central Avenue
D. Sonarpur
E. Howrah
Samples from the following nine sites were analysed for each
subject: dorsum of the tongue, lateral sides of the tongue, buccal
fold, hard palate, soft palate, labial gingiva and tonsils of soft tissue
surfaces, and supragingival and subgingival plaques from tooth
surfaces. samples were be collected by swab from the adjacent area
and was brought to the laboratory. Then samples were transported
to laboratory in ice-pack or within 4 hours of collection. Then the
samples were processed for bacteria and Yeast. Samples were
inoculated on the following media:
A. Mac Conkey agar with neutral red as pH indicator (Peptone,
Neutral red, agar agar, Lactose, Sodium taurocholate, deionized
water) for distinguish bacteria based on their lactosefermenting
properties into LF (Lactose-fermenting) and NLF
(Non-Lactose-fermenting) colonies.
B. Sabouraud’s dextrose agar plate (pH 5.6-6) (glucose -2gm,
Peptone 2 grams, Agar agar 2 grams, deionized water 100ml)
for fungi.
C. Robertson’s cooked meat medium (RCM) for culturing
anaerobes, made as per manufacturer’s instruction.
D. Blood agar plate for differentiate bacteria based on their
hemolytic properties.
E. Muller Hilton Agar for Antibiotic Susceptibility Test.
F. Nutrient Agar with 6.5% NaCl and Tellurite Agar for inoculation
of Enterococcus spp.
G. Egg Yolk Agar for inoculation of Bacillus spp.
Gram’s stain, Albert’s stain was done from samples directly to detect Gram positive or gram-negative bacteria, bacteria with metachromatic granules respectively. Sample inoculation and identification of Microorganisms.
The pure bacterial cultures were obtained by inoculating the sample on Blood agar media plates, MacConkey agar media plate, SDA tube and RCM broth. For this purpose, samples were streaked on different media and agar plates with the help of sterilized inoculating loops. The inoculated nutrient agar plates were then incubated for 18-24 h at 37 °C. After incubation the isolated bacterial colonies were picked from growth plates and quadrant streaking were done aseptically to new plates in order to obtain pure strains of bacterial culture. Four quadrants streaking were done by rotating the plates at 90° anticlockwise at four different areas of plate. This was done by dragging the culture across the agar with the help of sterilized inoculating loop from previously streaked area to new one. The plates were then incubated at 37 °C for 24h. After incubation isolated colonies were picked and cultured again to purify the samples [12,13].
Blood agar plate was used to grow fastidious microorganisms and to After culture, the plates were placed in incubator at 37 °C for a period of 18-24 hours for the development of colonies of bacteria, and after 24 hours the plate was taken out from incubator and different bacterial colonies were identified. Gram stain performed and after that the subculture of bacterial colony took place from the mixture. Biochemical tests like Indole, urease, motility, Citrate utilization, acid and H2S on TSI agar were also done from colonies of Gram-negative bacteria, and from Gram positive colonies, standard biochemicals like Catalase (with 3% H2O2), coagulase and oxidase were done [18,19].
Result
In the end of study, we were able to access the presence of pathogenic and normal bacteria in oral microbiome. Which are important of public health point.
How many samples tested
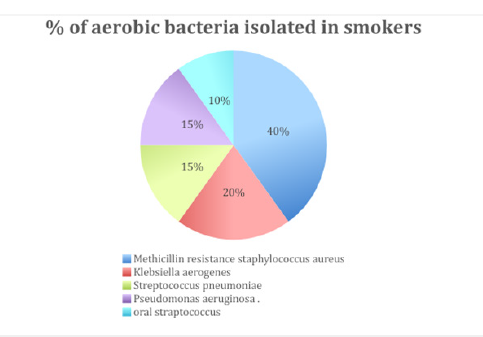
In this study we collected total 54 samples out of which 14 were taken from females and 40 from males. Out of 14 females 7 were smokers and 7 were non-smokers. Out of 40 males 20 were smokers and 20 were non-smokers. Out of 27 smokers 7 were occasional smokers while 20 were regular/chain smokers. Out of all the smokers, only one smokes bidi while others were cigarettesmokers. Out of all the smokers, 4 individuals were also consuming oral tobacco. The most common bacterial species we isolated overall, was Enterococcus spp. (44 isolates out of 54, 81.25%). Out of these 44 isolates, 34 were Enterococcus faecalis (77.27%) and 10 were Enterococcus non-faecalis (22.73%). Streptococcus pneumoniae was also isolated; 2 from smokers and one from non-smokers. In smokers most common pathogen found was MRSA (Methicillin Resistant Staphylococcus aureus) (50%) followed by Klebsiella aerogenes (20%) Streptococcus pneumoniae (15%) Pseudomonas aeruginosa (15%) and oral Streptococcus (10%) (Figures 1-3).
Figure 1:Gender distribution of Smokers vs nonsmokers.
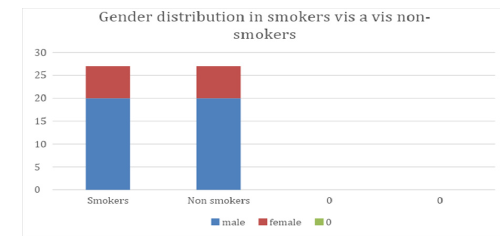
Figure 2:Age distribution of Smokers vs nonsmokers.
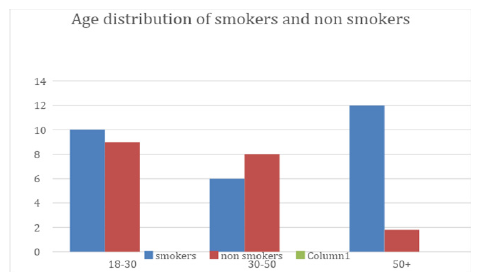
Figure 3:percentage of aerobic bacteria isolated from smokers.
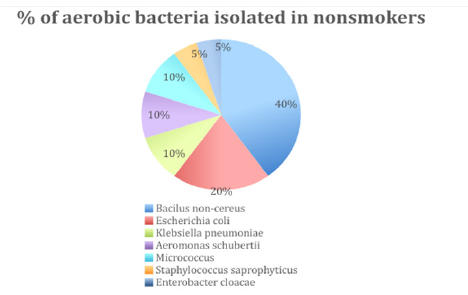
While in non-smokers the microbial flora isolated were more vivid as compared to smokers, Enterococcus faecalis and Enterococcus non- faecalis are most common as but no yeasts were found, the most common bacteria found in non-smokers were Bacillus non-cereus (40%) followed by coagulase negative Staphylococcus spp. (Staphylococcus saprophyticus) (5%). Apart from that the other common bacteria found were Enterobacter cloacae (5%), Aeromonas schubertii (10%), Escherichia coli (20%), Micrococcus (10%) and Klebsiella pneumoniae (10%). Prevotella, Veillonella and Streptococcus spp. were the predominant genera in the saliva of both groups. Although the overall composition and diversity of the microbiota were similar, Prevotella was significantly more abundant in salivary samples of current smokers compared to non-smokers (Figure 4).
Figure 4:percentage of aerobic bacteria isolated from Non-smokers.
Statistical calculation
MRSA was found more in smokers (14 out of 27) than Nonsmokers (0 out of 27). By taking 95% confidence interval and degree of freedom 1, with the help of chi-square test, this difference was found to be highly significant (p value <0.0001).
Discussion
Oral microbes are mostly Commensal, but they can also cause oral and periodontal disorders. They can vary according to the oral hygiene, and the smoking pattern of the host. At the end of the study, we were able to assess the burden of normal as well as pathogenic Microorganisms in smokers’ vis a vis nonsmoker. This type of study has not been done previously from this part of our country. Hence these findings were important from the public health viewpoint. The purpose of this study is to define the predominant Microbial flora of the healthy oral cavity by identifying and comparing the cultivable and the not-yet-cultivated bacterial species on different soft tissues and supra- and subgingival plaques. Our study was also helpful for finding the details about oral microbes. The microbiome of each organ is distinct and substantial variability occurs even within the oral cavity, partly due to spatial variations in the availability of Oxygen. The oral rinse samples used in this study are likely to overrepresent microbes present in the surface of the oral cavity as well as saliva, and less likely to include microbes from dental plaques or gingival crevicular fluid, which may explain the dominance of Firmicutes, as noted in buccal mucosa. Salivary microbiome has been reported to have higher richness and diversity compared with gingival plaque microbiome, and to be less susceptible to changes in periodontal conditions [17].
Conclusion
In conclusion, there is a distinctive bacterial flora in the healthy oral cavity which is different from that of smokers’ oral cavity. For example, many species specifically associated with periodontal disease, such as Porphyromonas gingivalis, Tannerella forsythia, and Treponema spp. were not detected in any samples tested. In addition, the bacterial flora commonly thought to be involved in dental caries and deep denticavities, represented by Streptococcus mutans, Lactobacillus spp., Bifidobacterium spp., and Atopobium spp., were not detected in supra- and subgingival plaques from clinically healthy teeth. More such studies are needed to delineate the exact difference of microbial flora between smokers and nonsmokers.
Source of Funding
None.
Conflict of Interest
There are no Conflicts of Interest.
References
- Ogba OM, Ewa JJ, Olorode OA, Mbah M (2017) Effect of tobacco smoking on oral microbial flora and the relationship with oral health in Calabar, Nigeria. Int J Bio Lab Sci 6: 1-5.
- Yamashita Y, Takeshita T (2017) The oral microbiome and human health. J Oral Sci 59(2): 201-206.
- Sedghi L, DiMassa V, Harrington A, Lynch SV, Kapila YL (2021) The oral microbiome: Role of key organisms and complex networks in oral health and disease. Periodontol 2000 87(1): 107-131.
- Costalonga M, Herzberg MC (2014) The oral microbiome and the immunobiology of periodontal disease and caries. Immunol Lett 162(2 Pt A): 22-38.
- Solbiati J, Frias Lopez J (2018) Metatranscriptome of the Oral Microbiome in Health and Disease. J Dent Res 97(5): 492-500.
- Yu G, Phillips S, Gail MH, Goedert JJ, Humphrys MS, et al. (2017) The effect of cigarette smoking on the oral and nasal microbiota. Microbiome 5(1): 3.
- Mason MR, Preshaw PM, Nagaraja HN, Dabdoub SM, Rahman A, et al. (2015) The subgingival microbiome of clinically healthy current and never smokers. ISME J 9(1): 268-272.
- Cole JR, Chai B, Farris RJ, Wang Q, Kulam SA, et al. (2005) The Ribosomal Database Project (RDP-II): Sequences and tools for high-throughput rRNA analysis. Nucleic Acids Res 33(Database issue): D294-D296.
- Camelo Castillo AJ, Mira A, Pico A, Nibali L, Henderson B, et al. (2015) Subgingival microbiota in health compared to periodontitis and the influence of smoking. Front Microbiol 6: 119.
- Moalic E, Gestalin A, Quinio D, Gest PE, Zerilli A, et al. (2001) The extent of oral fungal flora in 353 students and possible relationships with dental caries. Caries Res 35(2): 149-155.
- Fujimori I, Goto R, Kikushima K, Ogino J, Hisamatsu K, et al. (1995) Isolation of alpha-streptococci with inhibitory activity against pathogens, in the oral cavity and the effect of tobacco and gargling on oral flora. Kansenshogaku Zasshi 69(2): 133-138.
- Van Schooten FJ, Besaratinia A, De Flora S, D Agostini F, Izzotti A, et al. (2002) Effects of oral administration of N-acetyl-L-cysteine: a multi-biomarker study in smokers. Cancer Epidemiol Biomarkers Prev 2002 11(2): 167-175.
- Wirth R, Maróti G, Mihók R, Simon Fiala D, Antal M, et al. (2020) A case study of salivary microbiome in smokers and non-smokers in Hungary: analysis by shotgun metagenome sequencing. J Oral Microbiol 12(1): 1773067.
- Shankargouda P, Rao Roopa S, Sanketh DS, Amrutha N (2013) Microbial flora in oral disease. J Contemp Dent Pract 14(6): 1202-1208.
- Shankargouda P, Rao Roopa S, Sanketh DS, Amrutha N (2013) Oral microbial flora in health. World Journal of Dentistry 4(4): 262-266.
- Peter R, Emilia S, Uittamo J, Malcolm R, Riina R (2009) Novel method for sampling the microbiota from the oral mucosa. Clin Oral Investig 13(2):243-246.
- Chowdhry R, Singh N, Sahu DK, Tripathi RK, Mishra A, et al. (2018) 16S rRNA long-read sequencing of the granulation tissue from nonsmokers and smokers-severe chronic periodontitis patients. Biomed Res Int 2018: 4832912.
- Cintron F (1992) Initial processing, inoculation, and incubation of aerobic bacteriology specimens. In: Isenberg HD (Ed.), Clinical Microbiology Procedures Handbook, American Society for Microbiology, Washington DC, USA, 1: 1.4.1-.4.1.9.
- https://www.medcalc.org/calc/comparison_of_proportions.php
© 2023 Atul Raj and Sayan Bhattacharyya. This is an open access article distributed under the terms of the Creative Commons Attribution License , which permits unrestricted use, distribution, and build upon your work non-commercially.
 a Creative Commons Attribution 4.0 International License. Based on a work at www.crimsonpublishers.com.
Best viewed in
a Creative Commons Attribution 4.0 International License. Based on a work at www.crimsonpublishers.com.
Best viewed in 







.jpg)































 Editorial Board Registrations
Editorial Board Registrations Submit your Article
Submit your Article Refer a Friend
Refer a Friend Advertise With Us
Advertise With Us
.jpg)






.jpg)














.bmp)
.jpg)
.png)
.jpg)










.jpg)






.png)

.png)



.png)






