- Submissions

Full Text
Polymer Science: Peer Review Journal
Future Starch Analyses-Necessity for Higher Time and Local Resolution
Julia Compart and Joerg Fettke*
Biopolymer analytics, Institute of Biochemistry and Biology, University of Potsdam, Germany
*Corresponding author: Joerg Fettke, Biopolymer analytics, Institute of Biochemistry and Biology, University of Potsdam, Germany
Submission: January 16, 2021;Published: February 16, 2021

ISSN: 2770-6613 Volume1 Issue4
Abstract
Starch is chemically a relatively simple particle consisting of two homo-biopolymers, amylopectin and amylose, with only α-1,4 and α-1,6 linked glucosyl units. Starch is a natural storage carbohydrate in plants and algae and is an essential source for nutrition, animal feeding, as well as raw material for the industry. Despite increasing knowledge in the last decades, detailed structural information and understanding of its turn-over are largely lacking. Most data were generated using bulk experiments, what obviously now come to limitations for a deeper insight in starch metabolism. Here we focus on some unavoidable questions arising from existing data, related to starch biosynthesis, degradation, and structure, where these limitations strongly emerge.
Keywords: Starch; Starch structure; Starch metabolism; Starch biosynthesis; Starch breakdown
Introduction
Starch is the main energy provider in human nutrition as well as a raw material for industrial purposes and is mostly produced from cereal crops and tuber plants as e.g. potato. In most cases, higher plants form two types of starch, assimilatory (or transitory) starch and reserve (or storage) starch, which differ in quantity per plant, shape, size, metabolism, and biochemistry, but the internal structure of starch seems evolutionary conserved. Rising global temperature is causing major climatic challenges, particularly in terms of plant adaptation to changing environments. To sustain agricultural production, it becomes increasingly important to develop/identify and characterize plants that are able to adapt under such unfavorable growing conditions to secure crop production. This strongly demands for developing stress resistant varieties with improved starch characteristics for food as well as non-food production, such as in the pharmaceutical-, glue-, paper-, garments, textile-, bioplastic- and wood- industry. Therefore, a better knowledge of starch metabolism is needed.
Mini Review
So far, several parameters of starch are not understood, as e.g. structural composition,
the synthesis and degradation, regulation of the granule number, and the overall granule
morphology. It is not surprising that the number of proteins involved directly or indirectly
in starch metabolism is, and will further, increase. The analytics of starch in the moment is
often limited by the use of average data. Today our knowledge of starch reflected e.g. entire
tissues or organs. However, first indications reveal, that starch metabolism is altered at divers
leaf ages, development stages, or in different tissues [1-3]. Thus, characterization of single
tissues, cells, plastids, or starch granules are essentially missing, what is of high interest to
understand starch turn-over in more detail and requires the development of new starch
analytical techniques.
Clear limitations are related to the formation of starch granules. Here enzymatic/proteinic
as well as mechanistic data are widely lacking. Several mutants were described with altered
amount of starch granules per chloroplast in the model plant Arabidopsis. However, we are far
away from understanding the process of formation and also do not know all proteins involved.
The initiation of starch granules is highly complex, and several pathways are interconnected
and allow alternative possibilities [1,4-7]. Even the number of starch granules is not totally
fixed. Questions as, is the alteration related to different properties of the plastids, such as
metabolism and development, or are that statistically fluctuations arise. Similarly, it is not
understood why different sizes and morphologies of starch granules within one tissue, one
cell, or even one plastid occur.
Different pathways exist in plants for starch degradation,
especially when comparing the degradation of transitory and
storage starch. Also related to that, many questions are unanswered.
How the different starch particles get degraded simultaneously? Is
there an integration network that allows the detection of single
granule degradation? Is there a coordination of the degradation
in various plastids within one cell? Also here single cell, single
plastid, and single granule analyses are important. As example, for
various plants transitory starch degradation is connected with a
phosphorylation/dephosphorylation cycle on the starch granule
surface [8,9]. Thus, we do not understand, what defines the exact
phosphorylation position within an amylopectin molecule (Figure
1D), how deep in the surface of starch granules phosphorylation/
dephosphorylation occurs, are these events even distributed over
the surface and/or within an amylopectin molecule, exit differences
for starches from various plant species? These questions are also
interesting for industrial applications of starch, as phosphorylation
of starch is the only known natural covalent modification. This
together with the inner starch structure is important for the
physical properties of starches, which influences parameters as
swelling power, starch solubility, gelatinization, retrogradation,
syneresis, and rheological behavior.
Discussion
Figure 1: Open starch characteristics: the structure of amylopectin, the alternative phosphorylation positions within amylopectin, and the possible position of amylose. Alternative lamellae of crystalline and amorphous material (9nm repeat distance) formed by amylopectin molecules. Double helices (depicted as cylinders) are generated by short side chains of amylopectin that have vicinal intramolecular positions. These double helices can be organized as A-type and B-type allomorph (enlargement in illustration A). The orientation of the long chains of amylopectin molecules is the difference of the cluster model (A) in comparison to the building block backbone concept (B). In the later the long chains are oriented perpendicularly to the double helices and essentially parallel to the surface of the entire starch particle, whereas in the cluster model (A) they are oriented parallel to the double helices. The possible location of amylose (brown helices) is depicted in B. However, in principle also a mixture of both locations is possible. (D) The positions of phosphorylation within the amylopectin are depicted for C6- Phosphorylation. An even distribution of the phosphorylation, a random distribution of the phosphorylation positions, a phosphorylation only in close vicinity to the branching points, or phosphorylation close to the non-reducing end of a side chain is shown. However, also backbone chain phosphorylation is possible, but not illustrated.
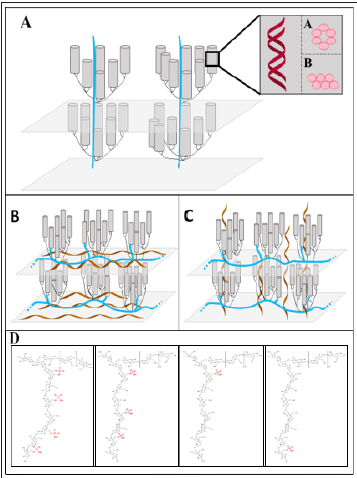
However, widely accepted structural models of both glucans forming starch, amylopectin and amylose are still lacking. Currently, two main models of amylopectin, the cluster model and the building block backbone concept exist, that describe the amylopectin structure (Figures 1A-1C). They differ in the position of the long chains. In the cluster model, the long chains have essentially the same orientation as the double helices and penetrate two or more layers of double helices. Whereas in the building block backbone concept, long chains are perpendicularly oriented to the double helices and form the backbone within each amorphous lamella [10-12]. However, also the precise location of amylose is obscure (Figures 1B & 1C). These models rely on average data and so our current knowledge is strongly limited. A higher time resolution of the analyses could show, whether starch granules during synthesis reveal altered inner structure compared with established granules. Moreover, this structural information would help to understand the initiation of starch granules and the enzymes/proteins that are involved. The inner starch structure seems to be evolutionary conserved. However, how this results in totally different starch granule morphologies is unknown. In planta flat, discoid, and spherical starch granules were reported. Also tiny granules that are smaller than 1μm and starches with more than 100μm can be detected. However, there are indications that the size and morphology are not only genetically determined, but also influenced by metabolic alterations of the plant [1,5,13]. Manipulation of starch granule morphology and size are important parameters for increasing starch yield and changing starch properties.
Conclusion
Overall, our knowledge related to structural, biosynthetic, and degradation aspects of starch metabolism has increased massively in the last decades, but we still lack basic understanding. Analyses will benefit from abandon average data and consider higher time resolution. Enlarging our insights will allow preparing starch-crops better adapted for the ongoing climatic changes.
Acknowledgment
This work was supported by the Deutsche Forschungsgemeinschaft DFG-FE 1030/2-1, 5-1, and 6-1.
References
- Malinova I, Mahlow S, Alseekh S, Orawetz T, Fernie AR, et al. (2014) Double knockout mutants of arabidopsis grown under normal conditions reveal that the plastidial phosphorylase isozyme participates in transitory starch metabolism. Plant Physiol 164(2): 907-921.
- Hammond JBW, Burton KS (1983) Leaf starch metabolism during the growth of pepper (Capsicum annuum) Plants. Plant Physiol 73(1): 61-65.
- MacNeill GJ, Mehrpouyan S, Minow MAA, Patterson JA, Tetlow IJ, et al. (2017) Starch as a source, starch as a sink: The bifunctional role of starch in carbon allocation. J Exp Bot 68(16): 4433- 4453.
- Malinova I, Qasim HM, Brust H, Fettke J (2018) Parameters of starch granule genesis in chloroplasts of arabidopsis thaliana. Front Plant Sci 9: 761-768.
- Malinova I, Alseekh S, Feil R, Fernie AR, Baumann O, et al. (2017) Starch synthase 4 and plastidal phosphorylase differentially affect starch granule number and morphology. Plant Physiol 174(1): 73-85.
- Seung D, Boudet J, Monroe J, Schreier TB, David LC, et al. (2017) Homologs of protein targeting to starch control starch granule initiation in arabidopsis Plant Cell 29(7): 1657-1677.
- Vandromme C, Spriet C, Dauvillee D, Courseaux A, Putaux JL, et al. (2019) PII1: A protein involved in starch initiation that determines granule number and size in Arabidopsis New Phytol 221(1): 356-370.
- Hejazi M, Fettke J, Kotting O, Zeeman SC, Steup M (2010) The laforin-like dual-specificity phosphatase SEX4 from arabidopsis hydrolyzes both C6- and C3-phosphate esters introduced by starch-related dikinases and thereby affects phase transition of alpha-glucans. Plant Physiol 152(2): 711-722.
- Steup M, Fettke J, Hejazi M (2011) The phosphorylation/dephosphorylation cycles in transitory starch metabolism. J Appl Glycosci 58: 162-163.
- Vamadevan V, Bertoft E (2015) Structure-function relationships of starch components. Starch 67(1-2): 55-68.
- Bertoft E (2015) Fine structure of amylopectin. In: Nakamura Y (Ed.), Starch Metabolism and Structure. Springer, Germany, p. 3-40.
- Hizukuri S (1986) Polymodal distribution of the chain lengths of amylopectins, and its significance. Carbohydr Res 147(2): 342-347.
- Malinova I, Fettke J (2017) Reduced starch granule number per chloroplast in the dpe2/phs1 mutant is dependent on initiation of starch degradation. PLoS One 12(11): e0187985.
© 2021 Joerg Fettke. This is an open access article distributed under the terms of the Creative Commons Attribution License , which permits unrestricted use, distribution, and build upon your work non-commercially.
 a Creative Commons Attribution 4.0 International License. Based on a work at www.crimsonpublishers.com.
Best viewed in
a Creative Commons Attribution 4.0 International License. Based on a work at www.crimsonpublishers.com.
Best viewed in 







.jpg)































 Editorial Board Registrations
Editorial Board Registrations Submit your Article
Submit your Article Refer a Friend
Refer a Friend Advertise With Us
Advertise With Us
.jpg)






.jpg)














.bmp)
.jpg)
.png)
.jpg)










.jpg)






.png)

.png)



.png)






