- Submissions

Full Text
Novel Approaches in Cancer Study
Modern Strategies in Cancer Study: Drug Repositioning in Colorectal Cancer Treatment
Anselmino LE, Malizia F and Menacho-Márquez M*
Instituto de Investigaciones para el Descubrimiento de Fármacos de Rosario (IIDEFAR, UNRCONICET), Universidad Nacional de Rosario, Rosario, Argentina
*Corresponding author:Menacho Márquez Mauricio, Research Institute for the Discovery of Drugs in Rosario, Argentina
Submission: April 16, 2019;Published: May 08, 2019

ISSN:2637-773XVolume2 Issue4
Abstract
Colorectal cancer is one of the most fatal cancers in the world because most of cases are diagnosed in advanced stages, when the development of resistance to chemotherapy is more frequent. To face this situation, new drugs and drug combinations would be necessary. Drug discovery is a very costly process; although efforts in drug discovery have been amplified in recent decades the success rate of approved FDA drugs continues being low. In response to this situation the repositioning of drugs proposes to delve into the genetic, epigenetic and metabolic differences to discover new targets and redirect drugs already approved to the treatment of other diseases and conditions. In this minireview we discuss the main avenues in which repositioning can occur and we cite some examples of repositioning drugs in colorectal cancer treatment that are being tested in clinical trials in the past years.
Keywords: Colorectal cancer; Drug repositioning; Clinical trials
Abbreviations: CRC: Colorectal Cancer; DCA: Dichloroacetate; DHEA: Dehidroepiandrosterona; EGFR: Epidermal Growth Factor Receptor; EMT: Epithelial-Mesenchymal Transition; FDA: Food and Drug Administration; GEO: Gene Expression Omnibus; NCBI: National Center for Biotechnology Information; 5-FU: 5-Fluorouracil
Introduction
Colorectal cancer (CRC) is the third most commonly diagnosed cancer in males and the second in females, with 1.8 million new cases and almost 861,000 deaths in 2018 [1]. CRC is often diagnosed at advanced stages, when the probability of development of distal or local recurrence due to chemotherapy resistance is more elevated [2,3]. The common protocol of CRC treatment consists in a primary surgical resection of the tumor, followed by radiotherapy and/or adjuvant chemotherapy. Since the 1950s, 5-fluorouracil (5-FU) remains the mainstay of chemotherapy [4,5]. In the recent years other drugs have been developed and used in combination with 5-FU such as oxaliplatin, irinotecan and capecitabine [6]. The use of new monoclonal antibodies such as Bevacizumab and Cetuximab has also allowed great advances in therapies [7]. However, almost half of patients with advanced CRC are resistant to chemotherapies based on 5-FU [8]. To counter this situation new strategies are being implemented; these include improved early diagnosis (down-staging), discovery of reliable predictive biomarkers and development of novel drugs/drug combinations.
The development of a new drug is a process that can take 10-15 years; the number of drugs approved by the FDA (Food and Drug Administration) has been declining since 1995 and investment in drug development has been gradually increasing, indicating that the cost of new drug development will continue to grow [9-11]. In this context, repurposing of drugs has emerged as an alternative to the classical pipeline of drug development and there has been increasing interest in analyzing anti-cancer activity of non-cancer drugs already FDA approved. The two main advantages of this strategy are the knowledge of the pharmacokinetics and toxicity of the drugs and their low costs and accessibility, since they are mostly generic [12]. In addition, repositioning of drugs is a strategy of special importance in low- and middle-income countries, where the conventional therapies are not economically accessible to the population in general due to their high costs [13,14].
Drug repositioning can take advantage of both computational and experimental methods. There are two main approaches in the selection of drugs to reposition: the first consists in propose a drug as possible anticarcinogenic based on prior information about it, such as the knowledge of its molecular target or the identification of its effects during the treatment of a clinical subpopulation that shows a lower incidence of certain disease. This approach is also called text mining-based since it is based on the literary and observational search of the mode of action and effect of the candidate drug and often the selection is follow by in vitro screening techniques. An example of this type is metformin, the postulation of this drug as a possible reposition drug arose as a result of a study of cancer incidence in patients with type II diabetes [15]. In this study it was observed that a group of diabetic patients treated chronically with metformin had a lower cancer incidence and cancer-related mortality. The drug was tested in vitro in cell lines of different types of cancer, including CRC, and due to favorable results also in animal models [16]. Preclinical studies determined that metformin affects cellular metabolism and suppresses oncogenic signaling pathways, such as receptor tyrosine kinase, PI3K/Akt, and mTOR pathway [17]. Metformin has been tested in combination with other compounds, showing synergistic action whit EGFR inhibitors and other repositioning drugs such as propranolol [18]. Currently, our group is testing the combination of metformin and propranolol on in vitro and in vivo models of CRC, where we observed a decrease in cell proliferation and tumor growth, and the inhibition of metastasis related events associated with an effect on epithelial-mesenchymal transitions through the modification of E-cadherin and B-catenin levels (unpublished data). Metformin is actually in phase II to be approved by the FDA for CRC treatment (Table 1).
Table 1:Some repositioning drugs in clinical trial stage for CRC treatment.
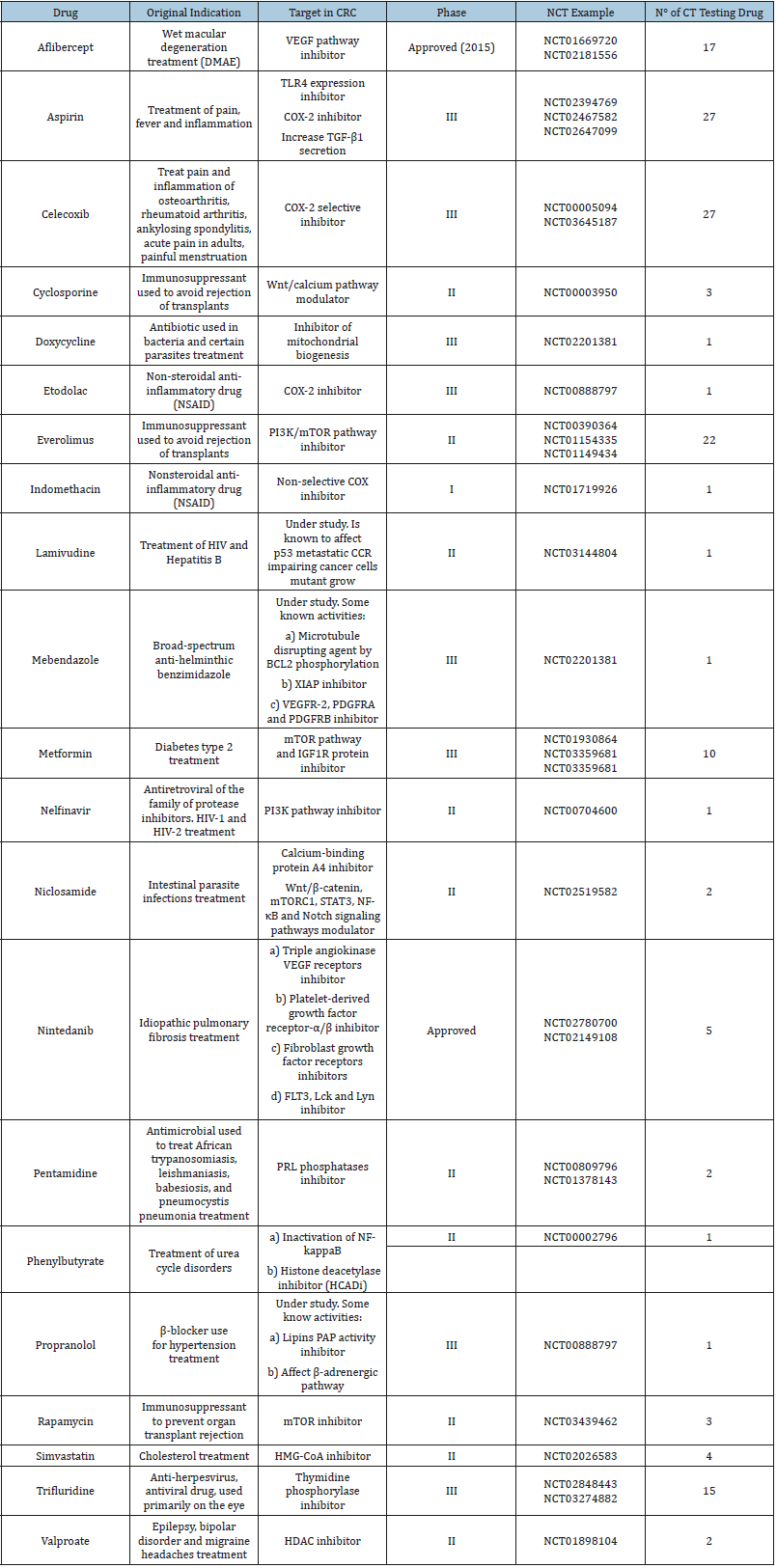
Within this same approach, other drugs were postulated to be repositioned based on their molecular target; an example is chloroquine, a drug used in the treatment of malaria. Chloroquine was proposed as a possible anticancer drug due to previous knowledge about its effects on parasite autophagy processes. It is known that chloroquine diffuses easily into lysosomes where is protonated and loses its ability to diffuse out of the vesicle. In malaria, chloroquine invades the lysosomes of the parasite and prevents the digestion of heme, killing the organism. In human cells, the accumulation of chloroquine in lysosomes produces the inhibition of enzymes such as phospholipase A2, hindering the breakdown of proteins and affecting cell signaling pathways [19]. The anticancer effects of chloroquine were corroborated in more than one type of cancer including CRC [20], leukemia [21] and breast cancer [22]. This drug is in phase II of preclinical studies for breast cancer treatment (NCT01446016) and in vitro stage for CRC treatment [23]. Other examples of repositioning drugs in CRC treatment whit this approach is dehidroepiandrosterona (DHEA) [24], orlistat [25], pantoprazole [26], DCA [27] and raloxifene [28], all these are in in vitro stage. A second alternative approach for the search of putative drugs to be repositioned takes advantage of bioinformatic methods. During the past decade, the development of new technologies to study gene expression and the advances in sequencing machinery, such as microarrays and the whole transcriptome shotgun sequencing (RNA-seq), have allowed the emergence of gene expression databases (most of which are freely available) putting a large amount of information at the disposal of the scientific community. Some databases, like Gene Expression Omnibus (GEO) at the National Center for Biotechnology Information (NCBI) [29], Array Express [30] and EMBL-EBI Expression Atlas [31] are repositories of a great variety of gene expression studies including genetic expression of cell lines to human tissues, all of them under the most varied experimental conditions. The analysis of this new information has led to great advances in the acquisition of biological and chemical data pertaining to alteration of pathways in diseases, alteration in protein structures, drug-target interactions, mechanism of drug actions, disease-specific and drug-induced gene expression signatures [32-34]. In silico drug repositioning is an approach that takes advantage of this situation to relocate drugs using computational strategies [34]. A large amount of drugs to be repositioned have been proposed based on this approach, some of them for CRC treatment are itraconazole [17], tanshinone IIA [35], and fluspirilene [36], all these still are in in vitro tests stage, while others are already being tested in humans CRC as rapamycin and celecoxib (Table 1).
Conclusion
Drug repositioning is becoming one of the fundamental pillars of personalized medicine; aim to improve the productivity of current drug discovery pipelines. The recent advances in next- generation sequencing technologies are enabling personalized genomic studies which in turn provide insight into the heterogeneity between patients within a disease and, in the case of cancer, even within a same type of tumor. The identification of new important molecular pathways and their respective pharmacological targets have potential therapeutic utility in drug repositioning. The knowledge derived these kinds of approaches will result particularly useful for those patients that have developed pharmacological resistance to conventional treatment and/or suffer from tumors with reduced therapeutically options, especially in low- and middle-income countries.
Acknowledgement
M.M-M. thanks ANPCyT- FONCyT (PICT 2014-2025, 2016- 1773) and CONICET for financial support.
Conflict of Interest
The authors declare that they have no competing interest.
References
- Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, et al. (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. A Cancer Journal for Clinicians 68(6): 394-424.
- André T, Boni C, Navarro M, Tabernero J, Hickish T, et al. (2009) Improved overall survival with oxaliplatin, fluorouracil, and leucovorin as adjuvant treatment in stage II or III colon cancer in the MOSAIC trial. J clin oncol 27(19): 3109-3116.
- Van der Jeught, K, Xu HC, Li YJ, Lu XB, Ji G, et al. (2018) Drug resistance and new therapies in colorectal cancer. World Journal of Gastroenterology 24(34): 3834-3848.
- Salonga, D, Danenberg KD, Johnson M, Metzger R, Groshen S, et al. (2000) Colorectal tumors responding to 5-fluorouracil have low gene expression levels of dihydropyrimidine dehydrogenase, thymidylate synthase and thymidine phosphorylase. Clinical Cancer Research 6(4): 1322-1327.
- Showalter, SL, Showalter TN, Witkiewicz A, Havens R, Kennedy EP, et al. (2008) Evaluating the drug-target relationship between thymidylate synthase expression and tumor response to 5-fluorouracil: Is it time to move forward? Cancer biology & therapy 7(7): 986-994.
- Yaffee P, Osipov A, Tan C, Tuli R, Hendifar A (2015) Review of systemic therapies for locally advanced and metastatic rectal cancer. Journal of gastrointestinal oncology 6(2): 185-200.
- Adler MJ, Dimitrov DS (2012) Therapeutic antibodies against cancer. Hematology/Oncology Clinics 26(3): 447-481.
- Douillard J, Cunningham D, Roth AD, Navarro M, James RD, et al. (2000) Irinotecan combined with fluorouracil compared with fluorouracil alone as first-line treatment for metastatic colorectal cancer: A multicentre randomised trial. The Lancet 355(9209): 1041-1047.
- Sertkaya A, Birkenbach A, Ayesha B, Eyraud J (2014) Examination of clinical trial costs and barriers for drug development. Report, US Department of Health and Human Services, Office of the Assistant Secretary for Planning and Evaluation, Washington, DC, USA, pp. 1-92.
- Kim IW, Hayoung J, Kim JH, Myeong G, Sangsoo K, et al. (2019) Computational drug repositioning for gastric cancer using reversal gene expression profiles. Scientific reports 9(1): p 2660.
- Xue H, Li J, Xie H, Wang Y (2018) Review of drug repositioning approaches and resources. International journal of biological sciences 14(10): 1232-1244.
- Pantziarka P, Bouche G, Meheus L, Sukhatme V, Sukhatme VP (2015) Repurposing drugs in your medicine cabinet: untapped opportunities for cancer therapy? Future oncology 11(2): 181-184.
- Leukemia EiCM (2013) The price of drugs for chronic myeloid leukemia (CML) is a reflection of the unsustainable prices of cancer drugs: From the perspective of a large group of CML experts. Blood 121(22): 4439- 4442.
- André N, Banavali S, Snihur Y, Pasquier E (2013) Has the time come for metronomics in low-income and middle-income countries? The Lancet Oncology 14(6): e239-e248.
- Evans JM, Donnelly LA, Emslie-Smith AM, Alessi DR, Morris AD (2005) Metformin and reduced risk of cancer in diabetic patients. BMJ 330(7503): 1304-1305.
- Kobiela J, Dobrzycka M, Jędrusik P, Kobiela P, Spychalski P, et al. (2018) Metformin and colorectal cancer-A systematic review. Experimental and Clinical Endocrinology & Diabetes.
- Saini N, Yang X (2018) Metformin as an anti-cancer agent: actions and mechanisms targeting cancer stem cells. Acta biochimica et biophysica Sinica 50(2): 133-143.
- Rico M, Baglioni M, Bondarenko M, Laluce NC, Rozados V, et al. (2017) Metformin and propranolol combination prevents cancer progression and metastasis in different breast cancer models. Oncotarget 8(2): 2874-2889.
- Baker NC, Ekins S, Williams AJ, Tropsha A (2018) A bibliometric review of drug repurposing. Drug discovery today 23(3): 661-672.
- Sasaki K, Tsuno NH, Sunami E, Tsurita G, Kawai K, et al. (2010) Chloroquine potentiates the anti-cancer effect of 5-fluorouracil on colon cancer cells. BMC cancer 10(1): 370.
- Chen X, Clark J, Wunderlich M, Fan C, Davis A, et al. (2017) Autophagy is dispensable for Kmt2a/Mll-Mllt3/Af9 AML maintenance and antileukemic effect of chloroquine. Autophagy 13(5): 955-966.
- Zhang Y, Cao Y, Sun X, Feng Y, Du Y, et al. (2017) Chloroquine (CQ) exerts anti-breast cancer through modulating microenvironment and inducing apoptosis. Int Immunopharmacol 42: 100-107.
- Verbaanderd C, Maes H, Schaaf MB, Sukhatme VP, Pantziarka P, et al. (2017) Repurposing drugs in oncology (ReDO)-chloroquine and hydroxychloroquine as anti-cancer agents. Ecancermedicalscience
- Osawa E, Nakajima A, Yoshida S, Omura M, Nagase H, et al. (2002) Chemoprevention of precursors to colon cancer by dehydroepiandrosterone (DHEA). Life Sci 70(22): 2623-2630.
- Kridel SJ, Axelrod F, Rozenkrantz N, Smith JW (2004) Orlistat is a novel inhibitor of fatty acid synthase with antitumor activity. Cancer Res 64(6): 2070-2075.
- Zeng X, Liu L, Zheng M, Sun H, Xiao J, et al. (2016) Pantoprazole, an FDAapproved proton-pump inhibitor, suppresses colorectal cancer growth by targeting T-cell-originated protein kinase. Oncotarget 7(16): 22460- 22473.
- Khan A, Andrews D, Blackburn AC (2016) Long-term stabilization of stage 4 colon cancer using sodium dichloroacetate therapy. World J Clin Cases 4(10): 336-343.
- Ahmed A, Marwa A, Ola N Sayed, Safaa H, Kista S (2017) 5-fluorouracil synergized with raloxifene and cytosine β-D- arabinofuranoside to combat colorectal cancers in vitro via controlling lipolysis. Journal of Pharmacology and Toxicology 12(1): 14-23.
- Clough E Barrett T (2016) The gene expression omnibus database. Methods Mol Biol 1418: 93-110.
- Brazma A, Parkinson H, Sarkans U, Shojatalab M, Vilo J, et al. (2003) Arrayexpress-a public repository for microarray gene expression data at the EBI. Nucleic Acids Res. 31(1): 68-71.
- Petryszak R, Burdett T, Fiorelli B, Fonseca NA, Gonzalez Porta M, et al. (2014) Expression atlas update-a database of gene and transcript expression from microarray-and sequencing-based functional genomics experiments. Nucleic Acids Res 42(Database issue): D926-32.
- Schenone M, Dančík V, Wagner BK, Clemons PA (2013) Target identification and mechanism of action in chemical biology and drug discovery. Nat Chem Biol 9(4): 232-240.
- Iorio F, Bosotti R, Scacheri E, Belcastro V, Mithbaokar P, et al. (2010) Discovery of drug mode of action and drug repositioning from transcriptional responses. Proc Natl Acad Sci U S A 107(33): 14621- 14626.
- Ma H, H Zhao (2013) Drug target inference through pathway analysis of genomics data. Adv Drug Deliv Rev 65(7): 966-972.
- Yang R, Y Lu, J Liu (2014) Identification of tanshinone IIA as a natural monoacylglycerol lipase inhibitor by combined in silico and in vitro approach. MedChemComm 5(10): 1528-1532.
- Patil SP, Pacitti MF, Gilroy KS, Ruggiero JC, Griffin JD, et al. (2015) Identification of antipsychotic drug fluspirilene as a potential p53- MDM2 inhibitor: A combined computational and experimental study. J Comput Aided Mol Des 29(2): 155-163.
© 2019 Menacho Márquez Mauricio. This is an open access article distributed under the terms of the Creative Commons Attribution License , which permits unrestricted use, distribution, and build upon your work non-commercially.
 a Creative Commons Attribution 4.0 International License. Based on a work at www.crimsonpublishers.com.
Best viewed in
a Creative Commons Attribution 4.0 International License. Based on a work at www.crimsonpublishers.com.
Best viewed in 







.jpg)































 Editorial Board Registrations
Editorial Board Registrations Submit your Article
Submit your Article Refer a Friend
Refer a Friend Advertise With Us
Advertise With Us
.jpg)






.jpg)














.bmp)
.jpg)
.png)
.jpg)










.jpg)






.png)

.png)



.png)






