- Submissions

Full Text
Modern Concepts & Developments in Agronomy
Search Advances on Resistance to Soybean Diseases Caused by Diaporthe-Phomopsis Complex: A Sustainable Strategy to Obtain Safe Food Products
Pioli RN1*, Peruzzo AM1, Hernández FE1, Cuba Amarilla M1, Pratta GR2, Chiesa MA3, Cairo CA3 and Morandi EN3
1Plant and Microbial Biodiversity Laboratory-Phytopathology area (BioVyM-FRE), Argentina
2Genetic and Improvement of Tomatoes (GMT), Argentina
3Plant Eco-Physiology Laboratory (LEFIVE), Argentina
*Corresponding author:Pioli RN, Plant and Microbial Biodiversity Laboratory-Phytopathology area (BioVyM-FRE), National University of Rosario, Argentina
Submission: June 14, 2023;Published: June 30, 2023

ISSN 2637-7659Volume13 Issue 1
Abstract
Diaporthe/Phomopsis (DP) is a fungal complex, hemi-biotroph that affects more than 900 hosts including cultivated and uncultivated species, native forests, fruits and weeds; they can also survive in seeds and stubble. The pathogenic plasticity of DP has been transmitted by seeds and expanded to different agro-ecosystems causing the inoculum to introduce a primary infection in disease-free batches of Argentina and diverse regions of the world. This situation prompted the necessity to characterize the pathogenic variability of Dp in the soybean-producing area of Argentina as a first step in developing effective strategies for incorporating Soybean Stem Canker (SSC) and Soybean Stem Blight (SSB) resistance into soybean germplasm. Collection of Diaporthe/Phomopsis isolates from different agro-environments has achieved some advances, such as: i) morphological determination and the first molecular validation into the identity of this fungal complex in Argentina, ii) detection and report that D. phaseolorum var. meridionalis (Dpm) and D. phaseolorum var. caulivora (Dpc) coexistence in the soybean producing area (32 to 34° SL), and iii) detection of variability in the virulence of different isolates of Dpm and Dpc when interacting with soybean genotypes carrying different resistance genes to SSC. This information explained the pathogenic results obtained, which showed that the soybean resistance genes that were effective against Dpm, were not for var. caulivora isolates [1]. To the four resistance genes to CTS-Dpm that were identified in soybean germplasm (Rdm1-4), a new gene was later determined as Rdm5 [2]. On the other hand, a major resistance gene of simple Mendelian inheritance to SSC-Dpc, named Rdc1, was detected and identified by classic genetic improvement and molecular assistance [3]; without discarding the possibility of another gene or genes that are associated with the assayed ones, which could of contributed to the resistance to SSC in soybean [4]. Respect to the Soybean Stem Blight (SSB) disease, caused by P. longicolla (Plo), it was possible to determine the first SSB resistance gene (Rpsb1) to SSB, carried by one of the resistant genotypes, without ruling out the possibility of carrying another associated genes [5]. These advances provide understanding about the effectiveness of the strategies applied and perspectives of plant improvement aimed at incorporating resistance to diseases in soybean.
Introduction
Diaporthe/Phomopsis (DP) is a fungal complex, hemi-biotroph that affects more than 900 hosts including cultivated and uncultivated species, native forests, fruits and weeds; they can also survive in seeds and stubble [6,7]. The trends and progress in biology, interactions with different hosts and bio-taxonomic adaptations since 1912 have already been summarized and published by several authors [1, 8-13]. The binomial specific generic: Diaporthe phaseolorum (Cooke& Ellis) Sacc. (Teleomorph, perfect sexual form, Dp) - Phomopsis phaseoli (Desmaz.) Sacc (anamorph, imperfect asexual form, Pp) and the independent species P. longicolla T.W. Hobbs (asexual form, imperfect o anamorph (Plo) of a teleomorph still unknown) [8,9] have been identified. The pathogenic plasticity of DP has been transmitted by seeds and expanded to different agro-ecosystems causing the inoculum to introduce a primary infection in disease-free batches of Argentina [14-16] and other countries of America [17-22], Europe, Asia and Africa [9,11,23]. Thus, DP has limited the production and quality of dry [16,24] and soybean fresh grain (edamame) [25] as crops and seeds of diverse native species of Argentina [26]. Dp complex interacting with Soybean causes two main pathologies: Soybean Stem Canker (SSC) and Soybean Stem and Pod Blight (SSB). SSC may be caused by D. phaseolorum var. meridionalis (Dpm) [2,8,27-30], and Dp var. caulivora (Dpc) [8,10,21,27,31-34]. Likewise, new reports have pointed out that D. aspalathi, previously cited as a pathogen of Aspalathus laminari by van Rensburg [13], was reported as a pathogen of soybean by Bruner et al. [35]. These results, according to taxonomic and biological concepts [7,8,36], would allow defining such D. aspalathi isolates as D. phaseolorum f. sp. aspalathi, a novel pathogenic variant to the soybean crop and belongs to a sub-specific taxon of D. phaseolorum. This conceptual framework introduces a different taxonomic perspective for the assigned denomination as an independent species of Diaporthe. Whilst SSB disease is caused by D. phaseolorum var. sojae (Dps) associated to P. longicolla (Plo), producing also seed decay in the reproductive stages of a soybean crop [7,8,11,23] (Figure 1-4).
Figure 1:Soybean stem canker by Diaporthe phaseolorum var. meridionalis (Dpm).
a) internal symptom.
b) external symptom (canker) covered by pycnidia (anamorph signs) of Dpm.
c) solitary perithecia (left) and perithecium with basal asco-carpus (teleomorph fruit body) (right).
d) asci and ascospores emerged from perithecia;
e) colonies of Dpm.
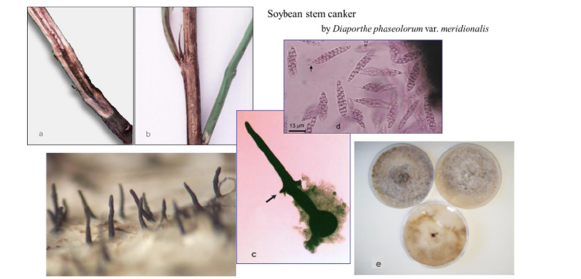
Figure2:Soybean stem canker by Diaporthe phaseolorum var. caulivora (Dpc).
a) symptoms in the field.
b) undefined cankers in stems (symptoms).
c) color and mycelial texture of Dpc colonies (left) and pycnidia (anamorph fruit body) scattered in the colonies
(right).
d) bundles of perithecia (teleomorph fruit body) associated to crop residues.
e) perithecia grouped under the epidermal cell layer;
f) perithecia grouped and forming vessels.
g) asci and ascospores emerged from perithecia of Dpc.

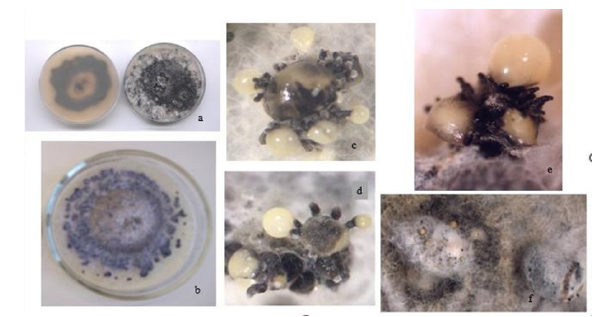
Figure 3:Soybean stem and pod blight by Diaporthe phaseolorum var. sojae (Dps).
a) stems and pods with blight symptoms and covered of pycnidia (anamorph fruit body).
b) color and mycelial texture of Dps colonies showing stromatic areas scattered in the Petri dishes.
c) colonies of Dps isolated from infected seeds.
d) d and e seeds covered with mycelium and mature pycnidia with drops or gutules of alpha and beta conidia.
e) f- perithecia of Dps formed on symptomatic stems.
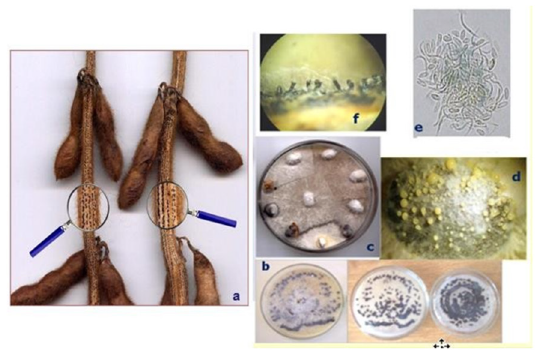
Figure 4:Soybean stem and pod blight by P. longicolla (Plo).
a) color and mycelial texture of Plo colonies (above).
b) stromatic areas scattered in the colony (bellow).
c) c-e typical bundles of pycnidia (anamorph fruit body) of Plo, showings their beaks delivering alpha conidia
as drops or gutules.
d) f-colony of P. longicolla obtained from infected seeds with the fungus.
Characterization of Diaporthe /Phomopsis - Glycine Max Interactions
The situation prompted the necessity to characterize the
pathogenic variability within Diaporthe/Phomopsis (DP) in the
soybean-producing area of Argentina as a first step for developing
effective strategies to incorporate of SSC and SSB resistance into
the soybean germplasm. A collection of DP isolates obtained from
different agro-environments allowed the first advances [10,37] to
be related to the following aspects:
i) the morphological determination and first molecular
validation of the identity of this fungal complex in Argentina,
ii) detection and report of Dpm and Dpc coexistence in the
soybean producing area (32 to 34° SL) [4,24], and
iii) detection of variability in the virulence of different Dpm
and Dpc isolates when interacting with soybean genotypes
carrying different SSC resistance genes, demonstrating genetic
variability in both varieties [29,38,39].
Inheritance of Resistance Genes to SSC Caused by Dpm
Analysis of the inheritance of resistance genes to SSC, during the 1980s and 1990s in the USA, determined the existence of four independent and single-inherited dominant genes. These non-allelic genes were named: Rdc1 and Rdc2, present in the cultivar (cv.) Tracy M [40-43]; Rdc3 and Rdc4, present in the cvs. Crockett and Dowling, respectively [40]. The Rdc4 gene was also identified in cv. Hutcheson [41]. However, according to the bibliographic revisions [1,9,11], the selection and incorporation of CTS resistance genes was carried out based on the inoculation of isolates of greater virulence obtained from different states in the southern USA (responsible for southern canker by Dpm) [10,41- 43]. This information explained the pathogenic results obtained, which showed that the soybean resistance genes that were effective against Dpm, were not for var. caulivora isolates. The above strongly suggested that genes for resistance to Dpm were different from genes for resistance to Dpc. Therefore, it was proposed to rename the genes Rdc1, Rdc2, Rdc3 and Rdc4, as Rdm1, Rdm2, Rdm3 and Rdm4, respectively; and among them, the higher level of resistance to SSC by Dpm was conferred individually by genes Rdm3 or Rdm4 [10].
Variability and Physiological Races in Dpm and Dpc Germplasm, Causal Agents of SSC
Pathogenicity trials also demonstrated the presence of physiological races in fungal germplasm of both varieties: Dpm [29] and Dpc causal agent of SSC [38,39]. On the other hand, the presence of two cultivars carrying the Rdm4 gene (Hutcheson and Dowling) allowed infer another virulence gene expressed in physiological races, which were virulent to Rdm4 only in the Dowling background. The fact that some Dpm isolates showed a different degree of compatibility when interacting with the same resistance gene in different backgrounds (e.g., Rdm4 gene in Dowling or in Hutcheson) suggested the existence of another gene or genes that are associated with the assayed ones, which could contribute to the resistance to SSC in soybean. Later, Chiesa et al. [2] identified a new gene, the Rdm5, linked to Rdm4 in Hutcheson cv., located at the Rdm4-5 locus; but Rdm (Rdm1-5) genes were not effective against SSC caused by Dpc [10]. Furthermore, the selection pressure given by the incorporation of Rdm genes for the resistance to SSC caused by Dpm in the soybean producing area, which promoted the expansion of the SSC disease caused by Dpc in Argentina [21,25,38]. Consequently, the SSC by Dpc gradually became one of the most important soybean diseases [38,39,44] because Rdc resistance genes had not been identified in the soybean germplasm and hence were not available for breeding programs.
Inheritance of Resistance Genes to SSC Caused by Dpc
In this context, the objective in this step of research was to identify and characterize the inheritance of Rdc genes for resistance to SSC-Dpc through classical Mendelian analysis, along with the assistance of specific molecular makers. Results obtained from specific and diverse interactions between Dpc16 and soybean genotypes demonstrated that G13 and G4 were, respectively, the most stable genotypes among the selected R and S parents. Thus, derived populations from F1, validated as hybrids and heterozygous by SNP, were selected to continue the analysis of the Rdc gene/s inheritance [44]. The chi-square goodness of fit test verified that phenotypic segregation of the complete F3 population adjusted to a 5:3 ratio (healthy resistant plants: dead susceptible plants) and phenotypic characterization of F2:3 families (Progeny Test) allowed to infer the genotypic ratio (1RR: 2Rr: 1rr) in the previous F2 individual population. Results obtained by classic genetic improvement and molecular assistance contributed to the detection and identification of a major resistance gene of simple Mendelian inheritance to SSC-Dpc, which was named Rdc1. Based on the updated bibliography revision, this was the first report on inheritance of Rdc resistance genes to SSC caused by Dpc [33].
Inheritance of Resistance Genes to SSC Caused by Dpc by Cuantitative Genetic Methods
Previous results showed that some populations from crosses of the same soybean genotype, carrying the Rdc1 gene, with other susceptible parents adjusted to Mendelian segregation after relaxing the required threshold of dead plants to qualify F2:3 families as phenotypically resistant [44]. Thus, the need arose to apply quantitative genetic methods that would allow validating the results obtained from the Mendelian genetics approach, detecting variations from eventual modifying genes in order to define strategies for the improvement in populations that are generated. The results validated G13 as a carrier of the Rdc1 gene, whose presence showed medium to high heritability. Further, in a population derived from G13 at least four segregating genes were detected for the incidence parameter of SSC by Dpc [4,44,45].
Inheritance of Resistance Genes to SSB Caused by Plo
Soybean Stem Blight (SSB) is caused by P. longicolla (Plo) and the binomial D. phaselorum var. sojae (Dps) (P. phaseoli var. sojae) [7]. In recent years, some soybean genotypes carrying Rpsd genes that confer resistance to soybean seed decay, caused by the same agents that cause SSB, have been reported. However, it was not known whether the Rpsd genes were also effective for Soybean Stem Disease (SSB). Therefore, in this case, the objective was the characterization of various soybean genotypes, carriers of Rpsd genes and others of interest, in order to evaluate their behavior against local strains of Phomopsis (Pps and Plo) that caused SSB, detecting eventual resistance genes (Rpsb) and determining their mode of inheritance. For this, effective crosses were obtained between differential genotypes (R and S) at SSB [5], and their respective F1 (molecularly validated by SNP) in order to continue the breeding program with segregating stable F2 and F3 populations. Through inoculations, the reaction against SSB of the parents, the F1, F2 individuals and the F3 plants distributed in the F2:3 family (Progeny Tests) were characterized. Through the observed phenotypic ratios, it was possible to infer the expected genotypic ratios in the F2 parents, allowing the identification of the first SSB resistance gene (Rpsb1) to SSB, carried by one of the resistant genotypes, without ruling out the possibility of carrying other associated genes [46-48].
These advances provided an understanding about the effectiveness of the strategies applied and perspectives of plant improvement aimed at incorporating resistance to diseases in soybean crops.
References
- Pioli R, Morandi E, Martínez M, Lucca M, Tozzini A, et al. (2003) Morphologic, molecular and pathogenic characterization of phaseolorum variability in Argentina. Phytopathology 93(2): 136-146.
- Chiesa MA, Pioli RN, Morandi EN (2009) Specific resistance to soybean stem canker conferred by the Rdm4 locus. Plant Pathology 58(69): 1032-1039.
- Peruzzo AM, Hernández FE, Pratta GR, Ploper LD, Pioli RN (2019) Identification and inheritance of an Rdc gene conferring resistance to soybean stem canker (Diaporthe phaseolorum caulivora). Eur J Plant Pathol 1180(154): 1179-1184.
- Cuba Amarilla Mario (2022) Identificación mediante enfoques cuantitativos de genes de resistencia al cancro del tallo (Diaporthe phaseolorum caulivora) en la base genética de soja. Tesis Maestría Genética Vegetal. Fac Cs Agrarias UN Rosario, Argentina.
- Hernández FE (2022) Búsqueda e identificación de genes de resistencia a Phomopsis longicolla en germoplasma de soja. Tesis Doctoral en Cs. Agrarias. Fac. Ciencias Agrarias UN Rosario, Argentina.
- Dissanayake AJ, Phillips AJL, Hyde KD, Yan JY, Li XH (2017) The current status of species in Diaporthe. Mycosphere 8(5): 1106-1156.
- Hernández FE, Peruzzo AM, Pratta GR, Pioli RN (2020) Identification of pathogenic diversity in Phomopsis sp. causing stem blight in soybeans (Glycine max) by molecular markers. Revista Agrociencia Colpos 54(3): 313-326.
- Grijalba PE (2011) Variabilidad morfológica, genética y patogénica de Diaporthe phaseolorum caulivora causante del cancro del tallo de la soja en la provincia de Buenos Aires. MS Thesis. Universidad Nacional de Mar del Plata. Balcarce, Argentina.
- Pioli RN, Morandi EN (2002b) Diaporthe phaseolorum meridionalis. Global Crop Protection Compendium. In: Lesley Mcgillivray (Ed.), CABI, Published by CD ROM versions, CAB Intern. Wallingford, UK, ISSN-1365-9065, p. 20.
- Pioli RN, Morandi EN (2005) phaseolorum var. sojae. Global Plant Protection Compendium. In: Lesley Mcgillivray (Ed.), CABI, Published by CD ROM, Wallingford, UK, ISSN-1365-9065, p.33.
- Pioli Rosanna N, Gosparini Carlos O, Ferri Mónica, Morandi Eligio N (2005/06) First report on pathogenic variability in the interaction Glycine max - phaseolorum var. caulivora in Argentina. (Abs. 101) USA. ISSN 0327 9545. Biocell 30(2): 404.
- Stewart S (2015) Caracterización del agente causal del cancro del tallo de la soja en Uruguay. Agrociencia Uruguay 19(1): 69-76.
- van Rensburg JC, Lamprecht SC, Groenewald JZ, Castlebury LA, Crous PW (2006) Characterisation of Phomopsis spp. associated with die-back of rooibos (Aspalathus linearis) in South Africa. Stud Mycol 55: 65-74.
- Keeling BL (1984) Evidence of physiological specialization in Diaporthe phaseolorum caulivora. Journal of the Mississippi Academy of Science 29(S): 5.
- Pioli R, Gattuso S, Prado D, Borghi A (1997a) Outbreak of soybean canker caused by Diaporthe phaseolorum meridionalis in Santa Fe, Argentina. Plant Dis 81(10): 1215.
- Rupe JC, Sutton EA, Becton CM, Gbur EE (1999) Effect of soybean growth stage at the time of inoculation with Diaporthe phaseolorum meridionalis on stem canker development and yield. Plant Dis 83(6): 582-586.
- Keeling BL (1988) Influence of temperature on growth and pathogenicity of geographic isolates of Diaporthe phaseolorum caulivora. Plant Disease 72(3): 220-222.
- Kilen TC, Keeling BL, Hartwig EE (1985) Inheritance of reaction to stem canker in soybean. Crop Sci 25(1): 50-51.
- Santos JM, Vrandečić K, Ćosić J, Duvnjak T, Phillips AJL (2011) Resolving the Diaporthe species occurring on soybean in Croatia. Persoonia - Molecular Phylogeny and Evolution of Fungi 27(11): 9-19.
- Costamilan Leila MJ, Yorinori T, ÁM Rodrigues Almeida, Claudine Seixas, Eliseu Binneck, et al. (2008) First report of phaseolorum var. caulivora infecting soybean plants in Brazil. Tropical Plant Pathology 33(5): 381-385.
- Hawksworth D, Kirk P, Sutton B, Pegler D (1995) Dictionary of the fungi. CAB International, UK, p. 616.
- Tyler JM (1996) Characterization of stem canker resistance in ¨Hutcheson¨ soybean. Crop Sci 36(3): 591-593.
- Yorinori JT (1996) Cancro da haste da soja: Epidemiologia e controle. Circ. Técnica N 14. Embrapa, Londrina, Brazil.
- Pioli RN, Morandi EN, Bisaro V (2001) First report of SSC caused by DPC, in Argentina. Plant Dis 85(1): 95.
- Benavidez R, Pioli RN, Morandi EN (2010) Response of the edamame edible soybean germplasm to Diaporthe phaseolorum, causal agents of soybean stem canker, in Argentina. Tropical Plant Pathology 35(1): 1-11.
- Alzugaray C, Carnevale NJ, Salinas AR, Pioli RN (2007) Biotic and abiotic factors that affect the quality of Schinopsis balansae and Aspidosperma quebracho-blanco Schltdl. seeds. Rev Iberoamericana de Micología 24(2): 142-147.
- Peruzzo AM, Hernández FE, Pratta GR, Ploper LD, Pioli RN (2017) First report on the identification and inheritance of an Rdc gene conferring resistance to soybean stem canker (Diaporthe phaseolorum caulivora). In IV Jornada Uruguaya de Fitopatología - II Jornada Uruguaya de Protección Vegetal, Montevideo, Uruguay.
- Pioli RN, Morandi EN, Gosparini CO, Borghi AL (1999b) First report on pathogenic variability of different isolates of Diaporthe phaseolorum meridionalis on soybean in Argentina. Plant Dis 83(11): 1071.
- Pioli RN, Benavídez R, Morandi EN, Bodrero M (2000) Epidemiological study of diseases associated to soybean carpels and seeds, in Santa Fe, Argentina. Fitopatología 35: 111-118.
- Chiesa MA, Pioli RN, Cambursano MV, Morandi EN (2013) Analysis of the resistance to soybean stem canker conferred by different Rdm loci in specific plant-pathogen interactions. European J Plant Pathology 135(2): 351-362.
- Pioli RN, Morandi EN (2002a) D. phaseolorum caulivora. In: Mcgillivray L (Ed.), Global Crop Protection Compendium, CAB International, Wallingford, UK: ISSN-1365-9065, ISBN-0-85199-649-3.
- Morgan-Jones G (1989) The Diaporthe/Phomopsis complex: taxonomic considerations. In: Pascale AJ (Ed.), World Soybean Res Conference IV Proceedings, Orientación Gráfica Editora, Buenos Aires, Argentina, pp. 1699-1706.
- Pioli RN, Porfiri A, Incremona M, Díaz C, Morata M (1993) Determination of fungic diseases in soybean (Glycine max). Previous study to evaluation of management methods. Com Biol 11: 156.
- Vidic M, Jasnic S, Miladinovic J (1995) Pathogenicity of sojae and P. longicolla. Zastita-Bilja 46(3): 197-205.
- Bruna BB, Valéria Lopes, Chicowski AS, Jessica DB, Fernanda M, et al. (2018) Morphological and molecular characterization of Diaporthe (anamorph Phomopsis) complex and pathogenicity of Diaporthe aspalathi isolates causing stem canker in soybean. Eur J Plant Pathol (2018) 151: 1009-1025.
- Alexopoulos CJ, Mims CW, Blackwell M (1996) Introductory Mycology. 4th (edn), J Wiley & Sons Inc., New York, USA, p. 880.
- Pioli RN, Cairo CA, Martínez MC, Benavidez R, Bisaro V, et al. (2009) Morphological molecular and pathogenic studies of phaseolorum var. caulivora in Argentina. In World soybean Research Conf. VIII, Beijing, China, p. 157.
- Pioli RN (2006) Situación del complejo Diaporthe phomopsis en el cultivo de Soja de Argentina. 3º Congreso de soja del Mercosur. Mercosoja. 27-20 junio 2006, Rosario. Argentina. Compendio de Conferencias, Foros y Workshops, pp. 325-331.
- Ploper LD (1989) The Diaporthe/Phomopsis disease complex of soybean. In: Pascale (Ed.), Proceedings. World Soybean Research Conference. Buenos Aires, Argentina, pp. 1695-1699.
- Bowers G, Ngeleka K, Smith O (1993) Inheritance of SSC resistance in Crockett and Dowling. Crop Sci 33: 67-70.
- Udayanga D, Xingzhong L, McKenzie EHC, Chukeatirote E, Ali HAB, et al. (2011) The genus Phomopsis, biology, applications, species concepts and names of common pathogens. Fungal Diversity 50: 189-225.
- Kilen T, Hartwig E (1987) Identification of single genes controlling SSC resistance. Crop Sci 27: 220-222.
- Lago ME (2010) Etiology and epidemiological aspects of soybean stem cáncer in the center and southeast of Buenos Aires. National University of Mar del Plata. Faculty of Agricultural Sciences, Argentina, p. 87.
- Peruzzo AM (2018) Búsqueda e identificación de genes de resistencia a la cancrosis del tallo de soja causada por Diaporthe phaseolorum caulivora. Tesis Doctoral en Cs. Agrarias. Fac Cs Agrarias. UN Rosario, Argentina.
- Cuba Mario, Alejandra Peruzzo, Guillermo Pratta, Rosanna Pioli (2021) Estimación del número de genes para resistencia a CTS (phaseolorum var. caulivora) en poblaciones segregantes. XVII Cong. Latinoamericano Genética.
- Ivancovich A (1992) Enfermedades del tallo de la soja: Carpeta de Producció Pergamino EE INTA, p. 11.
- Fernández FA, Hanlin RT (1996) Morphological and RAPD analysis of Diaporthe phaseolorum from soybean. Mycologia 88: 425-440.
- Hernández FE, Pioli RN, Peruzzo AM, Formento AN, Pratta GR (2015) Morphological and molecular characterization of a collection of isolates of Phomopsis longicolla (unknown teleomorph: Diaporthales) from the temperate and subtropical region of Argentina. Tropical Biology Journal 63(3): 871-888.
© 2023 Pioli RN. This is an open access article distributed under the terms of the Creative Commons Attribution License , which permits unrestricted use, distribution, and build upon your work non-commercially.
 a Creative Commons Attribution 4.0 International License. Based on a work at www.crimsonpublishers.com.
Best viewed in
a Creative Commons Attribution 4.0 International License. Based on a work at www.crimsonpublishers.com.
Best viewed in 







.jpg)































 Editorial Board Registrations
Editorial Board Registrations Submit your Article
Submit your Article Refer a Friend
Refer a Friend Advertise With Us
Advertise With Us
.jpg)






.jpg)














.bmp)
.jpg)
.png)
.jpg)










.jpg)






.png)

.png)



.png)






