- Submissions

Full Text
Environmental Analysis & Ecology Studies
Over Expression of Jatropha’s Dehydrin Jcdhn-2 Enhances Tolerance to Water Stress in Rice Plants
Samar A Omar1, Nabil I Elsheery2, Abdelnaser A Elzaawely2,Wacław Strobel3 and Hazem MKalaji3*
1Genetic Department, Tanta University, Egypt
2Agricultural Botany Department, Tanta University, Egypt
3Institute of Technology and Life Sciences, Poland
*Corresponding author: Hazem M Kalaji,Department of Plant Physiology, Faculty of Agriculture and Biology, Warsaw University of Life Sciences, Warsaw, Poland
Submission: June 06, 2018; Published: July 13, 2018

ISSN 2578-0336 Volume3 Issue3
Abstract
Jatropha curcas’sdehydrin (JC-DHNS) has been previously shown to play a role during natural dehydration process associated with maturation of Jatropha curcasseed. In this study, we generated transformed rice plant (tp) over expressing JC-DHNSgene and examined the role of over expressed gene in improving the drought tolerance. The tp plants showed a stronger growth under water stress condition induced by addition 20% of PEG 6000. It also showed an enhanced water stress tolerance as indicated by growth parameters included, fresh and dry weight, chlorophyll content, maximum quantum yield, actual quantum yield of photosystem II and non-photochemical quenching. Also, tp plants showed higher membrane stability under drought comparing with non-transformed plant (wt) as indicated through determination of membrane electrolyte leakage, the values of malondialdehyde and, hydrogen peroxide content as indicator for oxidation level. The tp plant had higher content of osmoregulators substances such as proline, free amino acids and total soluble sugar. The tp plant showed higher content of enzyme activity such as and superoxide dismutase catalase and ascorbate peroxidase compared with wt. Our results clearly showed that tp rice plant with JC-DHNS better coped with drought stress due increasing photosynthetic efficiency and antioxidant enzymes activity.
Keywords: Dehydrine; Over expression; Rice; Drought tolerance
Introduction
It’s well documented that drought is one of the greatest severe environmental stresses limiting the growth and yield of plants In Egypt, rice is cultivated as a summer crop in about one million faddan and is one of the main water consuming crops. The continuous flooding is the only method for irrigation and it consumes about 20% of the total water resources [1]. By the year 2025, with increasing the Egyptian population, it will be required to produce about 60% more rice than what is currently produced to meet the food needs of a growing population [2]. Few decades later, water availability will the limit factor of rice production in Egypt. Also, the regular global climate changes lead to the challenge of the world to produce enough food required to the increased population [3]. Improving genotypes productivity under water deficit condition is a great challenge for rice breeders to face water limitation problem. Drought tolerant plants have adopted various strategies to cope with environmental fluctuations through number of physiological, biochemical and molecular modifications and produced an array of proteins as a part of a global stress response to protect the cell metabolism [4-7]. Late embryogenesis abundant (LEA) encoding genes have been identified in numerous plant species in response to cellular dehydration and are, therefore, suggested to play a role in a biotic stress tolerance [7-9]. Dehydrins (DHNs) are classified as group 2 of LEA proteins [8,9] produced during late embryogenesis or drought, low-temperature, salinity, and ABA [4,7,10]. Several studies found that accumulation of dehydrin transcripts or proteins is associated with tolerance to freezing, drought, low temperature and salinity [11-15]. Recently, there are many direct experimental evidences correlating higher expression of DHNs and protection from osmotic stress. Arabidopsis plants over expressed a dehydrin fusion protein were found to have better survival to low temperature [16]. Also, transgenic tobacco expressing citrus dehydrin protein has been shown to give increased tolerance to low temperatures [17]. Over expression of wheat dehydrin DHN-5 improves tolerance to salt and osmotic stress in [18]. It has been suggested that the short amphipathic K segments of dehydrin polypeptides interact with solvent exposed hydrophobic patches on proteins undergoing partial denaturation and thus inhibit protein aggregate [10].
Jatropha curcas’s dehydrin (JC-DHN-5) has been identified and classified as typeY2SK2 according to the YSK shorthand for structural classification of DHNs [7]. Jc-DHN2 is 441bp long open reading frame and 156 amino acid residues with a predicted PI of 7.09 and special activity of β-N-acetyl hexosamini dases of glycoside hydrolase super family. Our previous data showed that JcDHN-2 transcript level showed a sharp increase in expression during the dehydration process occurred during seed maturation which supports the protection role of this kind of proteins against the harmful effects of dehydration. The aim of our study is to generate transgenic rice plants expressing JC-DHN-2 protein to investigate the contribution of this gene to improve the ability of plant to resist against drought stress. This investigation included the evaluation of some physiological changes accompanied such as growth parameters, photosynthetic activity, membrane stability measurements, osmolytes content and antioxidant enzymes activities for both transgenic (tp) and non-transgenic (wt) plants under both control and drought induced with PEG (20%) conditions.
Materials and Methods
Construction of a binary vector with JC-DHN2
figure 1: Creation of pCam-Dhn2. A- nucleotide seqJcDhn2 ORF (Accession No. KF113882( isolated from Jatropha curcas germinated seeds using JcDhn2 primers; B: the JcDhn2 was inserted between 35Spromotor and polya denylation signal 3’NOS termination in kpnI site of binary vector Pcambia1301-35SN to create the transformation vector Pcam-Dhn2; C: PCR screening of transformed plantlet using JcDhn2–ORF- specific primers. A specific PCR product of 490 bp was detected in tp and absent in wt.
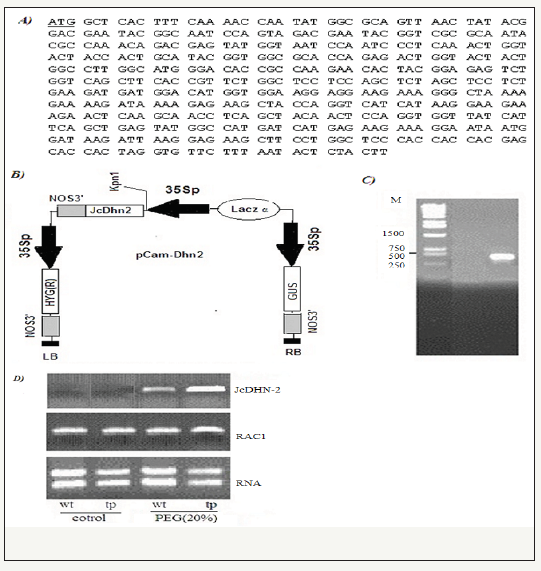
The full length JcDhn-2 open reading frame (ORF) (Acsession No KF113882(previously characterized by Omar et al. [7] was amplified from Jatropha curcas seedling using specific forward primer (ATGCTCACTTTCAAAACCAAT) and reverse primer (AAGTAGAGTATTAAAGAACAC). KpnI restriction site (GGTAC/C) were added to the JcDhn2 ORF and cloned into the KpnI site of the binary vector pCambia1301-35SN (kindly supported by -Prof Huang Jirong, shanghai institute of plant physiology and Ecology, shanghai, China) downstream the 35S promoter and upstream of poly adenenylation signal 3’NOS. The vector contains the hygromycin resistance gene hpt as a selectable marker between the 35S promoter and terminator. This vector was called pCam-Dhn2 (Figure 1). pCam-Dhn2 was mobilized into the Agrobacterium tumefaciens strains EHA105.
Rice calli production
Rice calli were generated from mature seed scutella of japonica rice cultivar (Oriza sativa L. Giza177) on a cullus induction medium (NB-AS medium containing 2,4-D-Dichlorophenoxyacetic acid (100mg/ml), casein enzymatic hydrolysate (1mg/ml), M-inositol (0.1mg/ml) and sucrose (30mg/ml). Sub culturing in same medium was repeated until obtaining light-yellow calli. Healthy growing light-yellow fragile calli were used for transformation and growing wt plants.
Transformation of rice
Rice transformation was achieved by co-cultivation of prepared calli with Agrobacterium tumefaciens EHA105 containing pCam- Dhn2 at 25-28˚C in dark for 3 days. The infected calli were selected using callus selection medium NB-THA containing: Timentin (100mglml), ampicillin (50mg/ml) and Hygromycin (50mg/ ml) for 2 weeks at 25-28˚C in dark. Calli were transferred to NBTH medium containing Timentin (100mg/ml) and Hygromycin B (50mg/ml) at 25-28˚C in dark for 10 days. Hygromycin resistant callus were transferred to the pre-regeneration medium containing benzylaminopurine BAP (1mg/ml), NAA (1mg/ml), ABA (12-15mg/ ml) and Hygromycin (50mg/ml) for 2 weeks Timentin (100mglml), ampicilin (50mg/ml) and Hygromycin (50mg/ml) for 2 weeks in dark at 25-28˚C. the resistant calli were transferred to regeneration medium containing 6-BAP(1mg/ml), NAA(1mg/ml),IAA(1mg/ml), KT(2mg/ml) and Hygromycin (50mg/ml) Timentin (100mg/ml), ampicilin (50mg/ml) and Hygromycin (50 mg/ml) for 2 weeks at 25-28˚C for 2-3 weeks. The regenerates were transferred to rootingmedium and incubated at 25-28˚C under light. Plants of 3-4 inches long are used for examining water stress tolerance.
PCR screening of transgenic plants
Non-transformed plants (wt) and transgenic plants (tp) were screened for the presence of JcDHN-2 using primers corresponding to the 5’and 3’ ends of JcDhn-2 with Kpn1 restriction sites added. These primers were 5’-GGTACCATGCTCACTTTCAAAAC-3’and 5’-GGTACCAAGTAGAGTATTAAAG-3’. The PCR was carried out in 25ul reaction volume according the instruction supporting with GoTaq® Green master Mix, (Promega) with implication condition as follows: 94˚C for 30sec, 54˚C for 1 min followed by 72˚C for 1 min. this was repeated for 30 cycles. The PCR product was examined on 1.5% agarose gel stained with Ethidium bromide.
Growth condition and stress treatment
Rice plants of wt and tp at 3-4 inches long were cultivated in 15cm pots. The experiment was carried out in the incubator at 28-29˚C and 14hrs light. Nutrient solution and nutrient solution containing 20% PEG-6000 was used for irrigation daily for both control and stress treatment, respectively. The concentration of PEG was maintained daily by changing the nutrient solution.
Determination of fresh weight, dry weight and water content
Fresh weight (FW) of whole plant was scored for individual plants. Dry weight (DW) of individual plants was determined after oven drying at 105˚C for 3h. Water content (wc) of individual plants was calculated as the difference between fresh and dry weight on base of FW and expressed as gH2O/g FW.
Measurements of photosynthetic parameters
Chlorophyll was extracted from leaf fragment (4cm long) using 2ml of 80% acetone. After incubation at 4 ˚C for 48hrs, the absorbance at 470, 649 and 665 nm was measured using a spectrophotometer [19]. Chlorophyll fluorescence was measured in 30 min dark adapted leaves using hand held leaf fluorometer (FluorPen FP 100, Photon System Instruments, Czech Republic). The following fluorescence parameters: maximum quantum yield of PSII in dark-adapted state (Fv/Fm), effective quantum yield of PSII, and non-photochemical quenching (NPQ) were measured according to [20,21].
Determination of osmolytes
Proline content was measured according to Bates et al. [22] total soluble sugars was extracted and assessed as described in Dubois et al. [23] total free amino content was determined as described earlier by Rosed et al. [24].
Determination of H2O2
Hydrogen peroxide was extracted with cold acetone according to the procedure described in Patterson et al. [25]. The extract was reacted quantitatively with titanium tetrachloride and ammonia to produce a peroxide-Ti complex. The complex was collected through centrifugation and then dissolved in 2M sulfuric acid. The absorbance of the solution was measured at 415nm and H2O2
content was calculated according to a standard curve.Evaluation of lipid per oxidation product
Lipid peroxidation was evaluated by determining malon dial dehyde (MDA) content from 0.5g of plant tissue as originally described by Heath et al. [26], with slight modifications by Hendry et al. [27].
Determination of antioxidant enzymes activities
Superoxide dismutase (SOD; EC1.15.1.1) activity was measured according to Beauchamp et al. [28] as described in Donahue et al. [29]. Catalase (CAT; EC1.11.3.6) activity was assayed according to the method of Aebi 1983. Ascorbate peroxidase (APX; EC 1.11.1.11) activity was assayed according to Nakano et al. [30].
RNA extraction and semi-quantitative PCR
Total RNA was extracted from wt and tp using Invi Trap® Spin plant RNA kit. DNA se-treated RNA (5μg) samples were used to synthesis of first strand cDNA according to the protocol supported by GoScriptTM reverse transcription Kit using oligo (dT) primer. Two microliters of the first strand cDNA were used as a template for PCR amplification with JcDhn-2 specific primers. The cDNA samples were standardized on rice actin gene (RAC1) transcript amount (accession no. X16280) using gene specific primers (F-CATGCTATCCCTCGTCTCGACCT, RCGCACTTCATGATGGAGTTGTAT). Samples were denaturized at 94˚C for 5 min and then run for 28 cycles at 94˚C for 30s, 5˚C 1min, 72˚C for 1min and final extension of 3 min at 72˚C. The PCR product was examined on 1.5% agarose gel stained with Ethidium bromide.
Results
Generation and identification of transgenic Rice of JCDHN- 2
JcDHN-2 full-length ORF (Figure 1A-1D) cloned into the KpnI site of the binary vector pCambia1301-35SN (kindly supported by Prof Huang Jirong, shanghai institute of plant physiology and Ecology, shanghai, china) downstream of the 35S promoter and upstream of poly adenenylation signal 3’NOS to produce pCam-Dhn2 vector (Figure 1A-1C) After Agrobacterium-mediated transformation, cullus transformation were confirmed by resistance to hygromycin as selectable marker between the 35S promoter and terminator. PCR analysis of non-transformed plants (wt) and transgenic plants (tp) using primers corresponding to the 5’and 3’ ends of JcDhn-2 with Kpn1 restriction confirmed the transformation and avoided the amplification of other DHNs from rice. Existing of one specific band in PCR product with size of 490bp confirmed the transformation in tp comparing with wt plants (Figure 1A-1C). To examine the expression level of JcDHN-2 in transgenic plants sq- RT-PCR was performed using gene specific primer to compare wt with tp under control and stress condition. (Figure 1A-1C) showed that no expression for JcDHN-2 in wt plants under both control and stress condition. Expression level of JcDHN-2 increased in tp plants as a result to stress condition comparing with control condition as appear in increasing of band density and brighten on agarose gel (Figure 1A-1C).
Evaluation of drought tolerance in JC-DHN-2 transformed plants
To study the role of Jc DHN-2 in drought tolerance of tp, we conducted several analyses and investigated stress-tolerance of wt and tp plants.
figure 2: Fresh and dry weight of plants under control and PEG (20%) treatment.

Fresh and dry weight
Determination of fresh (FW) and dry (DW) weight indicated to the superiority of tp plants on wt plants under stress condition. Tp plants kept their Fw and Dw values at high level under both conditions, while wt plants affected severely under drought condition (Figure 2).
Monitoring the oxidative state and membrane stability
figure 3: Stress tolerance monitor: changes in Electrolyte leakage rate of membrane, H2O2 content and MDA content of wt and tp plants under control and stress treatments.
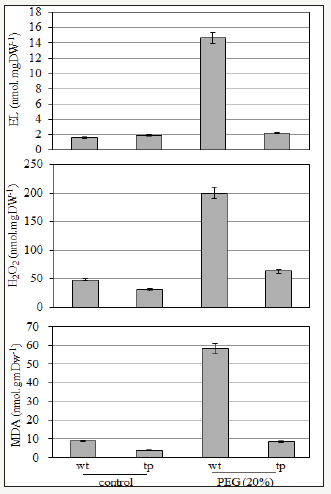
Tested plants showed that rate of EL, MDA and H2O2 content increased under stress conditions (Figure 3). Drought stress stimulates H2O2 production in wt compare with tp plants. The accumulation of H2O2 can cause oxidative stress in plant tissues and associated with increasing of MDA content as a direct indicator for membrane’s lipid oxidation. This severs effect of oxidation on membranes induce an increase in EL rate. Lower H2O2 content subsequently lower MDA content and EL rate were found in tp plants under drought indicated that over expression of JC-DHN-2 resulted in less ROS accumulation in tp plants. These results suggest a lack of capacity for protection from oxidative damage that occurred in wt under drought stress and reveal the great adaptation and membrane stability of tp plants to drought stress.
Analysis of photosynthesis pigments and their activity
figure 4: Effect of water stress on chlorophyl pigments and photosynthesis activity: changes on chlorophyl a,b , carotenoids contents and Fv/Fm on plants under control and PEG (20%) treatment.
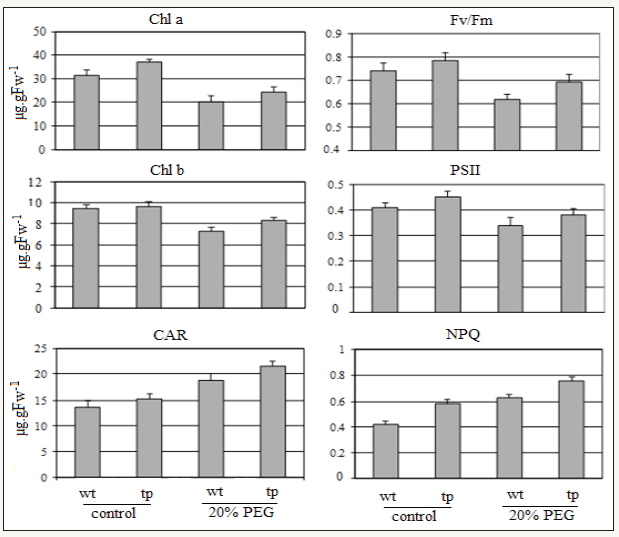
Determination of chlorophyll pigments such as a, b, and carotenoids and chlorophyll fluorescence parameters as Fv/Fm, PSII and NPQ showed increasing in their values in tp plants comparing with wt plants under control condition (Figure 4). Under stress condition, the contents of chlorophyll a, b, and carotenoids were greatly reduced in wt plants, whereas those values in tp plants were slightly reduced. These results clearly indicated that transgenic plants over expressing JC-DHN-2 had a greater tolerance to drought stress compared with wt plants.
Osmolytes compounds and antioxidant enzymes activities
Measurements of proline, FAA and TSS (Figure 5) showed that values of proline, TSS and FAA were higher in wt plants than tp plants under both conditions. Decreasing the values of proline, TSS and FAA in tp plants under drought conditions indicating that tp plants was less affected with drought than wt plants, where it reveal that these plants were protected against water loss. Determination of antioxidant enzymes (Figure 5) showed variation and differentiation between wt and tp plants. SOD enzyme showed similar activity in both wt and tp plants under control and also increased dramatically under drought stress in both of wt and tp. Although, CAT activity showed similar activity in both wt and tp plants under control condition, it showed great decrease in wt plants under drought stress while it increased slightly in tp plants under drought stress. APX activity showed higher values in tp plants comparing with wt plant under both control and drought conditions.
Discussion
JC-DHN-2 full-length ORF used to produce transgenic rice plants to cope with drought stress. In this study, transformed rice plants (tp) showed a great adaptation to drought stress treatment as shown by determination of membrane stability parameters (Figure 3) or growth and metabolism parameters (Figures 4,5). Numerous transgenic studies revealed a positive effect of DHNs genes expression on plant stress tolerance [31-34]. Dehydrins were may perform important protective functions in plant cells, such as protecting the structure and stabilizing the plasma membranes and raise the activity of stress-sensitive enzymes against drought stress [4,35]. Preservation of membrane integrity and stability under abiotic stress conditions is a main component of environmental stress tolerance in plants [36-38]. Transgenic plants over expressing DHNs showed less lipid peroxidation and leakage rate values [39,40]. The K-segments of DHNs can form amphiphilic α-helix, which may control the interaction of DHNs with lipid, plasma membranes and with hydrophobic sites of partially denatured proteins to prevent protein-protein aggregation of plasma membranes under drought stress [4,9]. Tobacco expressing spinach CAP85 and CAP160 DHNs revealed lower level of electrolyte after frost test which indicates a reduction of freezing injury in transformed plants [41,42]. Stimulating H2O2 production in wt plants under stress comparing with tp plant suggesting a lack of an enhanced capacity for protection from oxidative damage induced by drought stress in this wt. In this study, JcDHN-2 over expression prevents excessive accumulation of H2O2 under stress condition (Figure 3). Reactive oxygen species (ROS) scavenging function of DHNs was reported which can be mediated by direct interactions between the aa residue and the ROS species. Thus, it causes oxidation of the residue [43,44]. Moreover, DHNs can function as antioxidants (e.g., CuCOR15 and CuCOR19 in Citrus unshiu) [7,9,31]. Because of most DHNs function as molecular chaperons [9]. We could exclude that ROS level could be reduced by antioxidant enzymes protected by JCDHN- 2 expressed in pt plants under stress. Pattern of antioxidant enzymes activities showed good participation of SOD, CAT and APX in tp plants during stress treatment which can play role in ROS scavenging. Low activity of both CAT and APX in wt plants under drought stress caused losing of drought tolerance. CAT might be responsible for elimination of excess ROS during water stress [45,46] and remain more active for a greater duration of drought stress [47]. Accordingly, losing of CAT and APX activities in wt plants with increasing the activity of SOD resulted in accumulation of super oxide free radicals which cause severe effects on cell membrane and components. In our results, APX activity showed significant increase in tp plants under drought stress. APX and POD might be responsible for the fine modulation of ROS for signaling. Various stressful conditions of the environment have been shown to induce the activity of GPX [48], POD, APX, and GR [49,50] in tolerant species. It appears that there was an association between the higher antioxidant capacity and higher tolerance to drought stress in our transgenic rice which supports the idea of protection role of DHNs to antioxidant enzymes expressed in pt plants under water stress. Positive role of over expression of DHNs on relative water content and drought yield index as associated traits with drought tolerance occurred in set of Korean barley cultivars [12]. Also, the correlation between higher accumulation of DHNs transcript and drought tolerance was found in two differently tolerant cultivars of wheat [51]. DHNs contain high proportions of hydrophilic aa and change their conformation in response to the changes in their ambient micro-environment Hanin 2011 which led to changes in protein function [52,53].
figure 5: Changes in osmolytes contents and antioxidant avtivities: proline, FAA, TSS, SOD,CAT and APX for both wt and tp plant under control and PEG (20%) treatment under control and PEG(20%) treatment.

In conclusion, we here presented that rice transgenic plants over expressing JC-DHN-2 gene showed higher tolerance to water stress condition induced using 20% PEG 6000. Water stress tolerance in tp plants was accompanied with stability of membrane, increasing of photosynthetic parameters and good participation of some antioxidant enzymes. These results prove the protection role of DHNs under water stress condition.
Acknowledgment
This work is mainly supported by Tanta University Competitive Project Unit (TU-03-13-06), Egypt. Authors are grateful to Prof. Huang Jirong, shanghai institute of plant physiology and Ecology, Shanghai, China for his kindly support us with the binary vector (p Cambia1301-35SN). Authors are grateful to Prof. Zeng Fu Xu (Xishuangbanna Tropical Botanical Garden, Chinese Academy of Sciences, Kunming 650223, Yunnan, China) for his help.
References
- Aboulila AAM (2012) Molecular genetic studies on drought tolerance in the rice (Orzya sativa L.) using SSR DNA marker. Kafer El-Sheikh Egypt Fac of Agriculture, Egypt.
- Fageria N (2007) Yield physiology of rice. Journal of Plant Nutrition 30(6): 843-879.
- Shao HB, Liang ZS, Shao MA, Sun Q (2005) Dynamic changes of antioxidative enzymes of 10 wheat 4 genotypes at soil water deficits. Colloids and Surfaces B: Biointerfaces 42(3-4): 187-195.
- Allagulova C R, Gimalov F, Shakirova F, Vakhitov V (2003) The plant dehydrins: structure and putative functions. Biochemistry 68(9):945- 951.
- Ingram J, Bartels D (1996) The molecular basis of dehydration tolerance in plants. Annual Review of Plant Biology 47: 377-403.
- Omar SA, Fu QT, Chen MS, Wang GJ, Song SQ, et al. (2011) Identification and expression analysis of two small heat shock protein cDNAs from developing seeds of biodiesel feedstock plant Jatropha curcas. Plant Science 181(6): 632-637.
- Omar S, Elsheery N, Kalaji H, Ebrahim M, Pietkiewicz S, et al. (2013) Identification and differential expression of two dehydrin cDNAs during maturation of Jatropha curcas seeds. Biochemistry 78(5): 485-495.
- Saavedra L, Svensson J, Carballo V, Izmendi D, Welin B, et al. (2006) A dehydrin gene in physcomitrella patens is required for salt and osmotic stress tolerance. The Plant Journal 45(2): 237-249.
- Close T J (1997) Dehydrins: a commonalty in the response of plants to dehydration and low temperature. Physiologia Plantarum 100(2): 291- 296.
- Close T J (1996) Dehydrins: emergence of a biochemical role of a family of plant dehydration proteins. Physiologia Plantarum 97(4): 795-803.
- Choi DW, Close T (2000) A newly identified barley gene, Dhn12, encoding a YSK2 DHN, is located on chromosome 6H and has embryo-specific expression. Theoretical and Applied Genetics 100(8): 1274-1278.
- Park SY, Noh KJ, Yoo JH, Yu JW, Lee BW, et al. (2006) Rapid upregulation of Dehyrin3 and Dehydrin4 in response to dehydration is a characteristic of drought-tolerant genotypes in barley. Journal of Plant Biology 49(6): 455-462.
- Walia H, Wilson C, Wahid A, Condamine P, Cui X, et al. (2006) Expression analysis of barley (Hordeum vulgare L.) during salinity stress. Functional & Integrative Genomics 6(2): 143-156.
- Tommasini L, Svensson JT, Rodriguez EM, Wahid A, Malatrasi M, et al. (2008) Dehydrin gene expression provides an indicator of low temperature and drought stress: transcriptome-based analysis of barley (Hordeum vulgare L.). Functional & Integrative Genomics 8(4):387-405.
- Rorat T (2006) Plant dehydrins-tissue location, structure and function. Cellular & molecular biology letters 11(4): 536-556.
- Puhakainen T, Hess MW, Mäkelä P, Svensson J, Heino P, et al. (2004) Overexpression of multiple dehydrin genes enhances tolerance to freezing stress in Arabidopsis. Plant Molecular Biology 54(5):743-753.
- Hara M, Terashima S, Fukaya T, Kuboi T (2003) Enhancement of cold tolerance and inhibition of lipid peroxidation by citrus dehydrin in transgenic tobacco. Planta 217(2): 290-298.
- Brini F, Hanin M, Lumbreras V, Amara I, Khoudi H, et al. (2007) Overexpression of wheat dehydrin DHN-5 enhances tolerance to salt and osmotic stress in Arabidopsis thaliana. Plant Cell Reports 26(11): 2017-2026.
- Lichtenthaler HK,Wellburn AR (1983) Determinations of total carotenoids and chlorophylls a and b of leaf extracts in different solvents. Portland Limited 11(6): 591-592.
- Strasser RJ, Srivastava A, Tsimilli Michael M (2000) The fluorescence transient as a tool to characterize and screen photosynthetic samples. Probing photosynthesis: mechanisms, regulation and adaptation pp. 445-483.
- Kalaji HM, Schansker G, Ladle RJ, Goltsev V, Bosa K, et al. (2014) Frequently asked questions about in vivo chlorophyll fluorescence: practical issues. Photosynthesis Research 122(2): 121-158.
- Bates L, Waldren R, Teare I (1973) Rapid determination of free proline for water-stress studies. Plant and Soil 39(1): 205-207.
- Dubois M, Gilles KA, Hamilton JK, Rebers PT, Smith F (1956) Colorimetric method for determination of sugars and related substances. Analytical chemistry 28(3): 350-356.
- Rosed H (1957) Modified ninhydrin colorimetric analysis for acid nitrogen. Arch Biochem Biophys 67(1): 10-15.
- Patterson BD, MacRae EA, Ferguson IB (1984) Estimation of hydrogen peroxide in plant extracts using titanium (IV). Analytical Biochemistry 139(2): 487-492.
- Heath RL, Packer L (1968) Photoperoxidation in isolated chloroplasts: Kinetics and stoichiometry of fatty acid peroxidation. Archives of Biochemistry and Biophysics 125(1): 189-198.
- Hendry GA, Grime JP (1993) Methods in comparative plant ecology: a laboratory manual, Springer Science & Business Media, Germany
- Beauchamp C, Fridovich I (1971) Superoxide dismutase: improved assays and an assay applicable to acrylamide gels. Analytical biochemistry 44(1): 276-287.
- Donahue JL, Okpodu CM, Cramer CL, Grabau EA, Alscher RG (1997) Responses of antioxidants to paraquat in pea leaves relationships to resistance. Plant physiology 113(1): 249-257.
- Nakano Y, Asada K (1981) Hydrogen peroxide is scavenged by ascorbatespecific peroxidase in spinach chloroplasts. Plant and cell physiology 22(5): 867-880.
- Hara M, Terashima S, Kuboi T (2001) Characterization and cryoprotective activity of cold-responsive dehydrin from Citrus unshiu. Journal of Plant Physiology 158(10): 1333-1339.
- Hanin M, Brini F, Ebel C, Toda Y, Takeda S, et al. (2011) Plant dehydrins and stress tolerance: versatile proteins for complex mechanisms. Plant signaling & behavior 6(10): 1503-1509.
- Kumar M, Lee SC, Kim JY, Kim SJ, Kim SR (2014) Over-expression of dehydrin gene, OsDhn1, improves drought and salt stress tolerance through scavenging of reactive oxygen species in rice (Oryza sativa L.). Journal of Plant Biology 57(6): 383-393.
- Yin Z, Rorat T, Szabala BM, Ziółkowska A, Malepszy S (2006) Expression of a Solanum sogarandinum SK3-type dehydrin enhances cold tolerance in transgenic cucumber seedlings. Plant Science 170(6): 1164-1172.
- Sun X, Xi D, Feng H, Du J, Lei T, et al. (2009) The dual effects of salicylic acid on dehydrin accumulation in water-stressed barley seedlings. Russian Journal of Plant physiology 56(3): 348-354.
- Levitt J (1980) Responses of Plants to Environmental Stress, Volume 1: Chilling, Freezing, and High Temperature Stresses, Academic Press, USA.
- Filippou P, Antoniou C, Fotopoulos V (2011) Effect of drought and rewatering on the cellular status and antioxidant response of medicago truncatula plants. Plant Signaling & Behavior 6(2): 270-277.
- Elsheery NI,Cao KF (2008) Gas exchange, chlorophyll fluorescence, and osmotic adjustment in two mango cultivars under drought stress. Acta Physiologiae Plantarum 30(6): 769-777.
- Shekhawat UKS, Srinivas L, Ganapathi TR (2011) Musa DHN-1, a novel multiple stress-inducible SK3-type dehydrin gene, contributes affirmatively to drought-and salt-stress tolerance in banana. Planta 234(5): 915-932.
- Xing X, Liu Y, Kong X, Liu Y, Li D (2011) Overexpression of a maize dehydrin gene, ZmDHN2b, in tobacco enhances tolerance to low temperature. Plant growth regulation 65(1): 109-118.
- Campbell SA, Close TJ (1997) Dehydrins: genes, proteins and associations with phenotypic traits. The New Phytologist 137(1): 61-74.
- Kaye C, Neven L, Hofig A, Li QB, Haskell D, et al. (1998) Characterization of a gene for spinach CAP160 and expression of two spinach coldacclimation proteins in tobacco. Plant Physiology 116(4): 1367-1377.
- Hara M, Fujinaga M, Kuboi T (2005) Metal binding by citrus dehydrin with histidine-rich domains. Journal of experimental botany 56(420): 2695-2703.
- Helaly MN, El Hoseiny H, El Sheery NI, Rastogi A, Kalaji HM (2017) Regulation and physiological role of silicon in alleviating drought stress of mango. Plant physiology and biochemistry 118: 31-44.
- Helaly MN, El Sheery NI, El Hoseiny H, Rastogi A, Kalaji HM, et al. (2018) Impact of treated wastewater and salicylic acid on physiological performance, malformation and yield of two mango cultivars. Scientia Horticulturae 233: 159-177.
- Mittler R (2002) Oxidative stress, antioxidants and stress tolerance. Trends in Plant Science 7(9): 405-410.
- Rivero RM, Kojima M, Gepstein A, Sakakibara H, Mittler R, et al. (2007) Delayed leaf senescence induces extreme drought tolerance in a flowering plant. Proceedings of the National Academy of Sciences 104(49): 19631-19636.
- Kovalchuk I (2010) Multiple roles of radicals in plants reactive oxygen species and antioxidants in higher plants. pp. 31-44.
- Benešová M, Holá D, Fischer L, Jedelský PL, Hnilička F, et al. (2012) The physiology and proteomics of drought tolerance in maize: early stomatal closure as a cause of lower tolerance to short-term dehydration? PLoS One 7(6): e38017.
- Omar SA, Elsheery NI, Kalaji HM, Xu ZF, Song Quan S, et al. (2012) Dehydroascorbate reductase and glutathione reductase play an important role in scavenging hydrogen peroxide during natural and artificial dehydration of Jatropha curcas seeds. Journal of Plant Biology 55(6): 469-480.
- Labhilili M, Joudrier P, Gautier MF (1995) Characterization of cDNAs encoding triticum durum dehydrins and their expression patterns in cultivars that differ in drought tolerance. Plant Science 112(2): 219-230.
- Tompa P (2002) Intrinsically unstructured proteins. Trends in biochemical sciences 27(10): 527-533.
- Tompa P, Szasz C, Buday L (2005) Structural disorder throws new light on moonlighting. Trends in biochemical sciences 30(9): 484-489.
© 2018 Hazem M Kalaji. This is an open access article distributed under the terms of the Creative Commons Attribution License , which permits unrestricted use, distribution, and build upon your work non-commercially.
 a Creative Commons Attribution 4.0 International License. Based on a work at www.crimsonpublishers.com.
Best viewed in
a Creative Commons Attribution 4.0 International License. Based on a work at www.crimsonpublishers.com.
Best viewed in 







.jpg)































 Editorial Board Registrations
Editorial Board Registrations Submit your Article
Submit your Article Refer a Friend
Refer a Friend Advertise With Us
Advertise With Us
.jpg)






.jpg)














.bmp)
.jpg)
.png)
.jpg)










.jpg)






.png)

.png)



.png)






