- Submissions

Full Text
Approaches in Poultry, Dairy & Veterinary Sciences
Newcastle Disease Virus Respiratory and Digestive Viral Load Distribution Comparison
Orsi MA1,3*, Peixoto JM1, Zaroni MMH1, Camillo SCA1, Domingues CS1, Reischak D1, Gouvêa MV1, Freitas TRP2 and Arns CW3
1Federal Laboratory of Animal and Plant Health and Inspection-São Paulo (LFDA-SP), Campinas, São Paulo, Brazil
2Federal Laboratory of Animal and Plant Health and Inspection-Minas Gerais (LFDA-MG), Pedro Leopoldo, Minas Gerais, Brazil
1,3Department of Genetic, Evolution, Microbiology and Immunology, Institute of Biology, University of Campinas (UNICAMP), Campinas,SP, Brazil
*Corresponding author: Maria Angela Orsi, Department of Genetic, Evolution,Microbiology and Immunology, Institute of Biology, University of Campinas (UNICAMP), Campinas, SP, Brazil
Submission: January 04, 2021;Published: March 05, 2021

ISSN: 2576-9162 Volume8 Issue2
Abstract
Newcastle disease (ND) is one of most devastating disease of domestic and wild bird. Brazilian active and passive epidemiological surveillance have been implemented to avoid the Newcastle Disease Virus (NDV) infection risk. The present study was performed comparing the respiratory and digestive viral load distribution from apathogenic positive samples obtained from active surveillance and with the frequency through qPCR positive results from these swabs. The results show a higher load of NDV detected in respiratory samples and some swabs were positive only for cloaca demonstrating as both sampling as the molecular screening corroborate to improve the knowledge the NDV infection characteristics in the nature.
Keywords: Newcastle disease virus; Diagnosis; Active surveillance; qPCR
Introduction
Newcastle disease virus (NDV) belongs to Orthoavulavirus genus of the Paramyxoviridae family [1]. The NDV is responsible for a highly contagious and widespread avian disease that affects all bird species, and it is one of the most economically relevant diseases to the poultry industry. The active surveillance collecting samples from poultry slaughtered, migratory birds and imported genetic material and production with ≥10% mortality. The passive surveillance based on virus screening in suspicious samples. The samples collected for active surveillance were mainly cloacal and tracheal swabs, that arrived at laboratory in aliquots that varies in number (2,3 to 6 for each tissue) which were analyzed by reverse Real-time PCR (qPCR) to detect the NDV- M gene. The comparing the respiratory and digestive viral load distribution of apathogenic ND from positive samples and the frequency through qPCR positive results from these swabs were performed. The medium values of cycle threshold (Ct) obtained were also compared.
Material and Methods
Samples processing
The swabs have maintained in MEM and BHI culture mediums with antibiotics until centrifuged at 8,000rpm for 10min. An aliquot of 100μL of each supernatant from the 31 samples. The extraction of total nucleic acid using equipment Magna Pure LC. The reverse real-time PCR (qPCR) was carried on with AgPath ID™ One-Step kit at Applied Biosystems 7500 RT-PCR cycler. The protocol of the National Veterinary Services Laboratories (NVSL) [2] was applied. The primer sequence has consisted of 120 base pairs detected by fluorescence probe. The Ct was calculated automatically by the apparatus at the time that the amplification of viral cDNA. Analyzing the relationship between Ct and the proportion of positive results for each matrix (tracheal and cloacal swabs) noticed that this proportion decreases as the Ct value increases until it reaches its limit (Ct>40). Statistical analysis: The comparison of the Ct values for the paired samples since the samples have been processed simultaneously considering the null hypothesis is no difference between the results of PCR from different swabs. Hypothesis Ho: pt≤pc vs Ha: pt>pc.
Result
The median Ct found and its tolerance interval of 95% probability to the trachea was 36.89 (35.48 to 37.70) and the cloaca 36.76 (32.27 to 38.00) to fit relationship by probit analysis (SAS, 2000). However, considering that the two arrays are from the same record, there were a greater number of samples in positives tracheal than in cloacal swabs. It was confirmed by the comparison test of proportions [3], under the hypothesis Ho: pt ≤pc vs Ha: pt>pc, in which Ho was rejected (p=0.003), emphasizing the greater positivity in tracheal swabs than in the cloaca. The estimated difference between proportions and confidence intervals of 95% was 0.12 (0.03 to 0.22), showing that from respiratory system (tracheal) is most probable the detection of the virus. In the Figure 1 showing the difference between proportion of results in swabs of trachea and cloaca, by registration. The Figure 2. Proportion of positive results in swabs of trachea and cloaca on same samples, by registration.
Figure 1:Difference between proportion of positive results in swabs of trachea and cloaca, by registration.

pT= Proportion of Positive Results in Trachea Swab; pC=Proportion of Positive in Cloaca Swab
Figure 2:Proportion of positive results in swabs of trachea and cloaca on same samples, by registration.
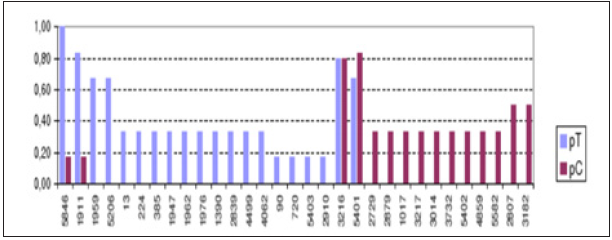
pT=Proportion of Positive Results in Trachea Swab; pC=Proportion of Positive in Cloaca Swab
Discussion
This study explores the distribution of respiratory and digestive viral loads in birds identified as infected with NDV in active surveillance by LDFA-SP. The results have demonstrated the detection of virus predominance in respiratory system (trachea). Molecular techniques allow the amplification of the viral genome directly from infected tissues, avoiding the need for isolation of the virus. However, its performance is hindered by the presence of PCR inhibitors in various organs and tissues, especially in the feces [4] which may explain the difference observed in the detections made from swabs. Gohm & Hofmann [5] had reported success in expanding a 182bp product, including the cleavage site, directly from infected chicken tissues. The main disadvantage is that no tissue was always positive during a time evolution study, implying the need to test a wide variety of tissues and organs.
Conclusion
These results have showed a higher load of NDV detected in respiratory tissue (trachea) and that, the higher frequency of positive results is in this tissue. Nevertheless, some swabs were positive only for cloaca. Such results demonstrate the importance of the type of sample collected and used for the molecular diagnosis of screening for ND, emphasizing the need for concomitant collection of both swabs to increase the chances of NDV detection.
References
- Amarasinghe GK, Ayllón MA, Bào Y, Basler CF, Basler CF, et al (2019) Taxonomy of the order Mononegavirales: update 2019. Arch Virol 164(4): 1967-1980.
- Wise MG, Suarez DL, Seal BS, Pedersen JC, Senne DA, et al. (2004) Development of a real-time reverse-transcription PCR for detection of newcastle disease virus RNA in clinical samples. J Clin Microbiol 42(1): 329-338.
- Snedecor GW & Cochran WG (1980) Statistical Methods. (7th edn), Iowa State University Press, UK, p. 507.
- Wilde J, Eiden J, Yolken R (1990) Removal of inhibitory substances from human fecal specimens for detection of group rotavirus by reverse transcriptase and polymerase chain reactions J Clin Microbiol 28(6): 1300-1307.
- Gohm DS, Thür B, Hofmann MA (2000) Detection of Newcastle disease virus in organs and faeces of experimentally infected chickens using RT-PCR. Avian Pathol 29(2): 143-152.
© 2021 Orsi MA. This is an open access article distributed under the terms of the Creative Commons Attribution License , which permits unrestricted use, distribution, and build upon your work non-commercially.
 a Creative Commons Attribution 4.0 International License. Based on a work at www.crimsonpublishers.com.
Best viewed in
a Creative Commons Attribution 4.0 International License. Based on a work at www.crimsonpublishers.com.
Best viewed in 







.jpg)






























 Editorial Board Registrations
Editorial Board Registrations Submit your Article
Submit your Article Refer a Friend
Refer a Friend Advertise With Us
Advertise With Us
.jpg)






.jpg)














.bmp)
.jpg)
.png)
.jpg)










.jpg)






.png)

.png)



.png)






